A Brief Introduction
This report is meant to provide an update on the status of the Big Y results within the Pike DNA Project, concentrating on developments that have transpired since the March 2018 update.As a quick reminder, the Big Y test analyses on the order of ten million nucleotide positions on the Y chromosome. One thing that this means is that with this single test it is possible to determine whether the person being tested is positive or negative for a multitude of SNPs (Single Nucleotide Polymorphisms) all at one time. Members of our project typically find that they are positive for around 950 to 1150 SNPs that have previously been identified, named, and incorporated in the overall Y-SNP Tree at Family Tree DNA.
But more than this, the test also detects new and previously undiscovered mutations. These are a key part of what we are hoping to find within our project, because these variants in the DNA of the Y-chromosome will help us to build Pike SNP trees that associate these mutations with different Pike family branches. When these variants are initially found they are identified only by their location on the chromosome. In time, those which are found to be present in multiple people are given SNP names, and may also become added into the overall Y-SNP Tree at Family Tree DNA. As one example, earlier this year such a variant was found at location 8141867 in the results of a member of our project's Group 6. When results for another member of this group revealed the same variant was present, it got named as the SNP BY50719. It has since been added to the overall Y-SNP Tree, where you can see it here. Other examples of SNPs that have similarly been identified and placed into the tree thanks to tests from within our project are BY165777 from Group 10 and BY24054 from Group 2. The SNP named YP5461 wasn't named as a result of Big Y test results, but determining its position in the tree was made possible by Big Y results from our Group 1.
In the various sections of this webpage, the Big Y test results from each of our project's genetic groups are analysed. In each case a SNP tree for the group is developed, based on the Big Y test results from that group and also taking into account known genealogical details when possible. One of the things that I'm particularly happy about is that in cases where there are insufficient historical records to build a connected family tree, we are now beginning to see just what our family tree truly looks like. This is something that will continue to happen as more members of our project undergo the Big Y test and more Pike-specific SNPs are discovered and then placed into the trees.
It should be understood that the nucleotide positions are not individually tested. That is, the Big Y test does not bundle together ten million individual tests. Rather, stretches of DNA are tested, often several times. The "reads" that cover each position then provide multiple glimpses of each position and whether or not it contains a variant that is different from the standard reference value for that position. Family Tree DNA rates each read as being of high/medium/low quality, and depending on this quality as well as how many reads were successful for each position they are able to identify nucleotides at which mutations are present. Sometimes different reads of a location yield different results, in which case Family Tree DNA will sometimes report a "no call" for the position. These variations in certainty can make analysing Big Y results tedious, since each reported variant ought to be manually double checked to determine whether it can be used with confidence. Examples will be shown in the discussions below. As for the reference values for each position, some information about how the human reference genome was established can be found in this recent post in Roberta Estes' "DNA Explained" blog, which also gives a description of how Family Tree DNA presents Big Y test results.
Note that in addition to the SNP results that are provided by the Big Y test, Big Y test results also automatically include details for over 500 STRs (Short Tandem Repeats), which is the type of marker that comprises the usual 111 marker results of our project. When ordering the Big Y test there is no need to order a separate upgrade in order to test any STRs that have not yet been determined. These 500+ STR values will also play a role in shaping our family trees, but there aren't yet many administrative comparison tools provided by Family Tree DNA for those beyond the first 111 of them, so I'm only beginning to manipulate the data from the STRs. See the discussion below regarding our project's Group 2 for some discussion about these STRs.
I encourage people to fully read this entire document, even if they are interested primarily in just one of our project's genetic clusters. Each of our groups offers something to help with learning and understanding Big Y results.
Something to notice when reading this document is that many of the deductions and conclusions that we are able to make are only possible because we are drawing upon the results from several people. My point here is that it takes a collective effort for us to be able to achieve these accomplishments. And having said that, I'll briefly say that more Big Y results from Pikes who have not yet done the test would very much be welcome.
Group 1
Within Group 1, which is our project's largest genetic cluster, we now have three Big Y test results, from Larry (kit number 129135), Alan (kit number 120594), and Roger (kit 70909), each of whom traces his ancestry back to John Pike who was born around 1572 in or near Whiteparish in Hampshire. In 1635 John, along with his children, left England and settled in Massachusetts.In terms of deeper haplogroup origins, our Group 1 falls within Y-DNA Haplogroup R1a, and more specifically within its L664 subgroup. Within the L664 subgroup is a sub-subgroup that is currently defined by the pair of markers named YP4102 and YP5464. This sub-subgroup contains not just our Group 1 cluster of Pikes but also non-Pikes, so we can be confident that these two SNPs arose prior to the Pike family. All three of Larry, Alan and Roger share a set of seven markers that the non-Pikes don't carry, and so we can find a branching point in the tree, with one side leading to non-Pikes and the other side leading to our Group 1 (as a brief side comment, in the previous BigY analysis done in March I drew a tree that showed a non-Pike who shared them... I'm not sure why now; I reckon that either I made a mistake in drawing the tree then, or maybe there's been a change in what is reported at FTDNA). Since Larry, Alan and Roger share these seven markers, these markers must pre-date (or first occurred within) their common ancestor John Pike, whose position in the tree is indicated with the oval bearing his name. Note that some of these seven markers might be universal within Group 1, but only by testing Pikes who do not descend from John will we be able to find out. Here's the current SNP Tree for Group 1, which will be followed by some further discussion about how it was determined.
FTDNA currently reports that Larry and Alan differ by three SNP mutations, Larry and Roger by five, and Alan and Roger by three. The variant named BY28714 is shown as a difference when comparing Larry to the other two (Larry is reported as not carrying it, and the other two as carrying it). Upon closer inspection using FTDNA's Y-Chromosome Browser it appears that BY28714 has produced low quality results for Alan and Roger. Interestingly, BY28714 also appears to no longer be represented in FTDNA SNP Tree, so possibly this marker has been deemed to not be consistently reliable. So despite being reported as positive for Alan and Roger, we'll omit it from this version of our Pike SNP Tree.
The remaining two mutations that FTDNA reports as differences between Larry and Alan consist of one that is named A2475 and another found at location 9000668 and not yet given a proper name Since 9000668 is reportedly present in Larry's results but not in those for Alan or Roger, we can deduce that this mutation must have arisen somewhere within Larry's lineage, subsequent to when it split away from each of Alan and Roger's lineages. Since we know how all three of these men descend from settler John, we can further say that this mutation at location 9000668 must have happened in one of six men, namely Larry or his father Francis or grandfather Willis or ... or great great great grandfather Zachariah who was born in 1755.
The situation with A2475 is less clear. Below you can see what is shown when we inspect it with FTDNA's Y-Chromosome Browser for each of Larry, Alan and Roger. We start with what the browser shows for Larry, for which we see that there is strong consistency at this marker's location (look at the values shown below the black arrow). As all but two of the reads of this position are the same as the reference value, it strongly appears that Larry is negative for this SNP.
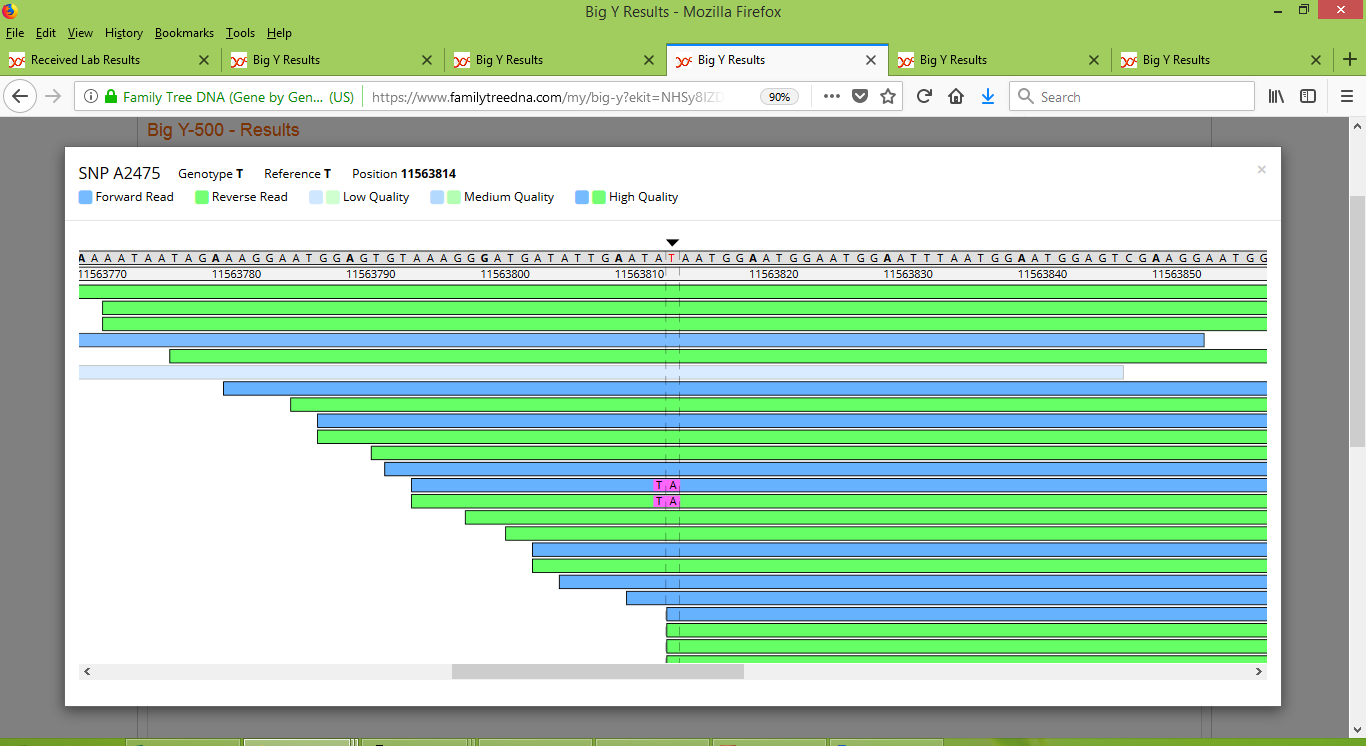
For Alan, we again see strong consistency, but this time the results are oppositve to those of Larry. Notice how Alan has a string of "A" results for the marker, whereas Larry's results are almost all the same as the reference value (which happens to be "T" for this location). So Alan strongly appears to be positive for this SNP.
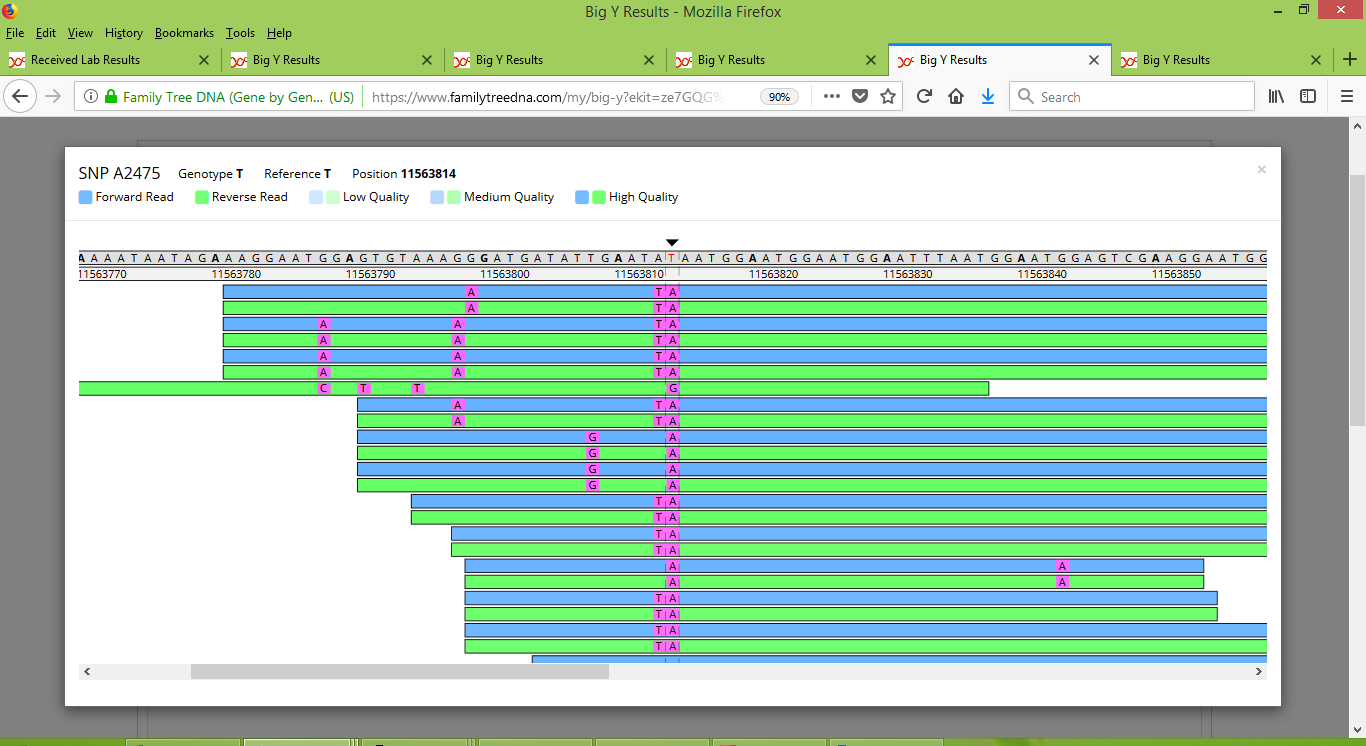
For Roger the results are about half the same as and half different from the reference value. FTDNA did not report a result for Roger for A2475, but instead reported it as a "no call". The ambiguity of his results explains why.
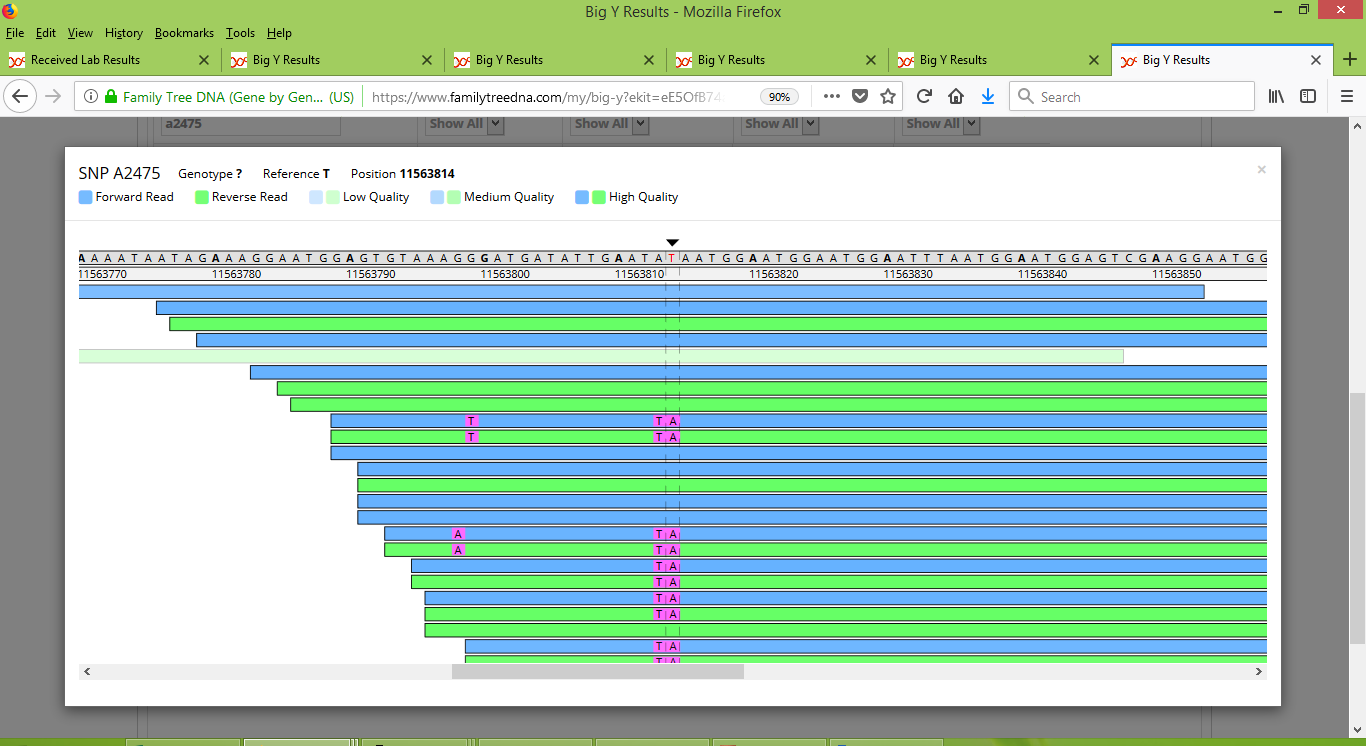
FTDNA reports that Larry is positive for A2475 and that Alan is negative. Using their two results alone, it would appear that the founder of our Group 1 was positive for A2475 but that Alan's line experienced a back-mutation at this location. However, given the ambiguous results for Roger, along with my understanding is that such backward mutations for SNPs are extremely rare, I contacted FTDNA to ask for some clarification. They advised that A2475 is a centromeric mutation that they don't consider to be of phylogenetic significance. To underscore their stance, they mentioned that about half of the Big Y results from individuals in haplogroup R1a have yielded a positive result for for A2475, which would imply that just under half have returned a negative value, which in turn shows that this SNP isn't reliable for tree-building purposes. Accordingly, A2475 has not been included in the SNP tree shown for Group 1. Marker A2475 also illustrates another point, namely that working with Big Y results can be tricky! This theme will recur when we look at the Big Y results for the other groups in our project.
Meanwhile, Roger has three not-yet-named variants at locations 13545985, 13547050 and 17359321 that are reported for neither Larry nor Alan. We conclude that they arose as mutations sometime along his lineage between him and his 8x great grandfather Robert Pike (i.e., along the part of his lineage that is not shared with Larry or Alan). Although these variants were discovered when his Big Y test was done about a year ago, they will remain unnamed and only identified by their location until such time as when somebody else is found to carry the same mutations.
Even though no further unnamed variants have currently been identified for Larry, Alan or Roger, that does not necessarily mean that no more will be found. Having watched some other people's results I've seen variants newly appear for them from time to time, usually about the time that another family member receives Big Y results. What seems to be happening is that some variants might have fallen short of some threshold to warrant being reported at first, but when additional results come along and provide additional supporting evidence, the variant then gets reported (and unnamed variants also tend to be given names at about this time too). We'll see some examples of this happening when we discuss the latest results for Group 2 below. Incidentally, this is also another way in which analysing Big Y test results can be tricky.
If I were able to wave a magic wand and select which members of Group 1 would be best to next have tested with the Big Y test, my highest priority would be to test somebody who does not descend from settler John. My thinking here is that testing somebody who is not closely related to the current Big Y testers would give us the best chance of sorting out the seven SNPs that are listed just above John in the SNP tree. What I mean by this is that some of these seven SNPs may be peculiar to just a portion of our Group 1 family tree, but right now we can't tell if that's the case or not. As for who to test, I actually think there are two priorities here: one is to test somebody who shares the value of 25 on the second of the STR markers (i.e., the second out of the usual 111 markers that most of us have tested), and also to test somebody who carries the value 24 on this marker.
Group 6
Within Group 6 we now have four Big Y test results available, whereas we only had two back in March. These four results are for a brother of Karen (kit number 61277), our project's co-administrator Stuart (48191), and most recently Russell (83604) and Gordon (398544). Stuart, Russell and Gordon are all known to descend from James Pike who was son of James Pike who settled in Massachusetts around the 1640s. Although the available records suggest that Karen likely descends from James too, there are some gaps in the records that have hindered making a firm conclusion.The Big Y test results that we have on hand now provide us with evidence that Karen does indeed descend from James. When we only had results from Karen and Stuart, Family Tree DNA reported that Karen (well, actually her brother) carried an unnamed variant at location 8141867 but Stuart did not carry it. At the time this merely indicated that Karen's line had acquired a mutation at some point after when it split from Stuart's. And when that split happened was uncertain.
To reach the conclusion that Karen descends from James, we first note that both Russell and Gordon also carry the mutation at location 8141867. And once it was found to be carried in multiple people, Family Tree DNA assigned a name to it. So instead of just talking about location 8141867, it can now be referred to as the SNP named BY50719. Since Russell, Gordon and Stuart each have traced their Pike lineages back to settler James, and in particular to his son James junior, we can work with this genealogical knowledge and isolate just when BY50719 arose. Russell and Gordon descend from James junior's son Nathaniel (born in 1685), whereas Stuart descends from James junior's son Samuel (born in 1690). Stuart doesn't carry BY50719, so it must have first occurred within a man who is an ancestor of both Russell and Gordon but who is not an ancestor of Stuart. James junior's son Nathaniel is such a man, as his son Samuel (born in 1730). Since Russell and Gordon descend from two different sons of this Samuel, we can therefore conclude that BY50719 must have arisen either with Nathaniel or his son Samuel.
When discussing Group 1 we saw that it is important to manually inspect each SNP with the FTDNA Y-Chromosome Browser. When we do this for BY50719, the results are plain and unambiguous. Stuart is clearly negative for BY50719, as we can see in the screenshot below from the Y-Chromosome Browser (as a side note, eight nucleotides to the right of BY50719, at location 8141875 Stuart has two spurious reads with "A" alleles).
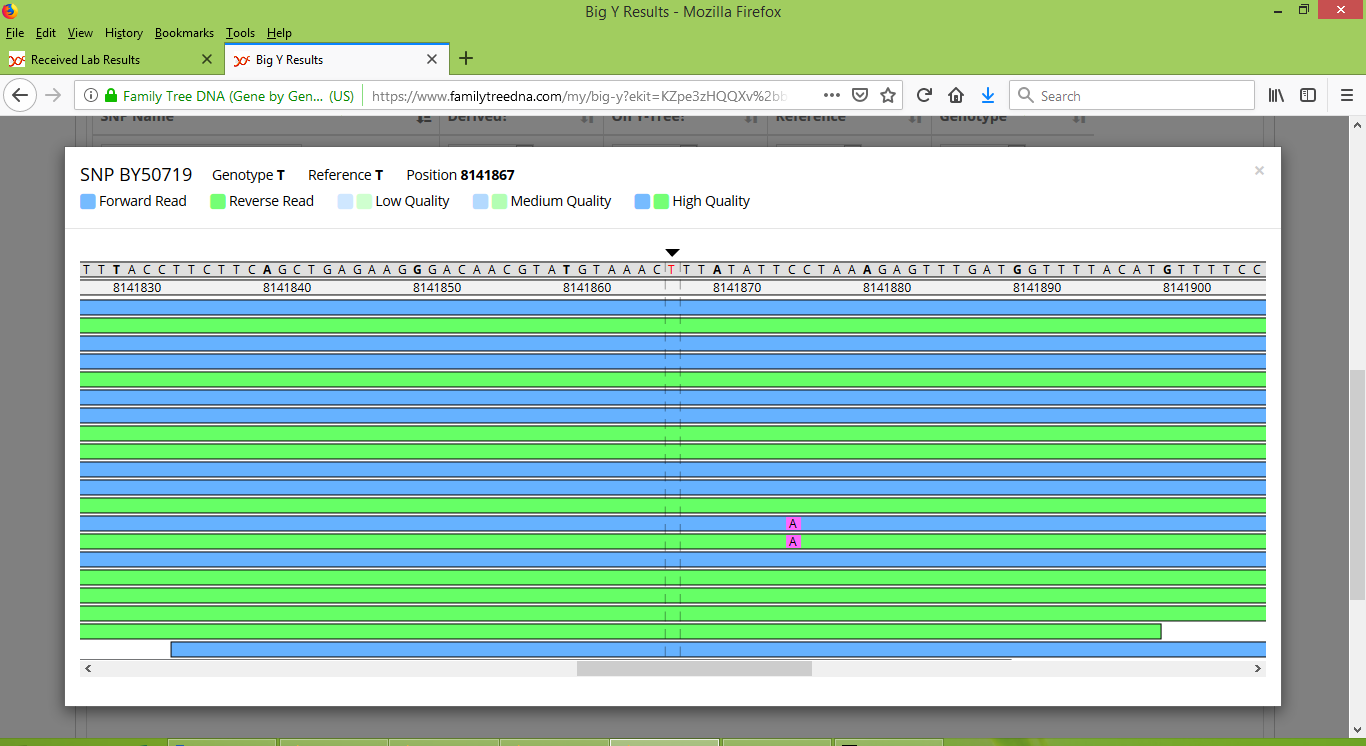
And Karen's brother is clearly positive for BY50719:
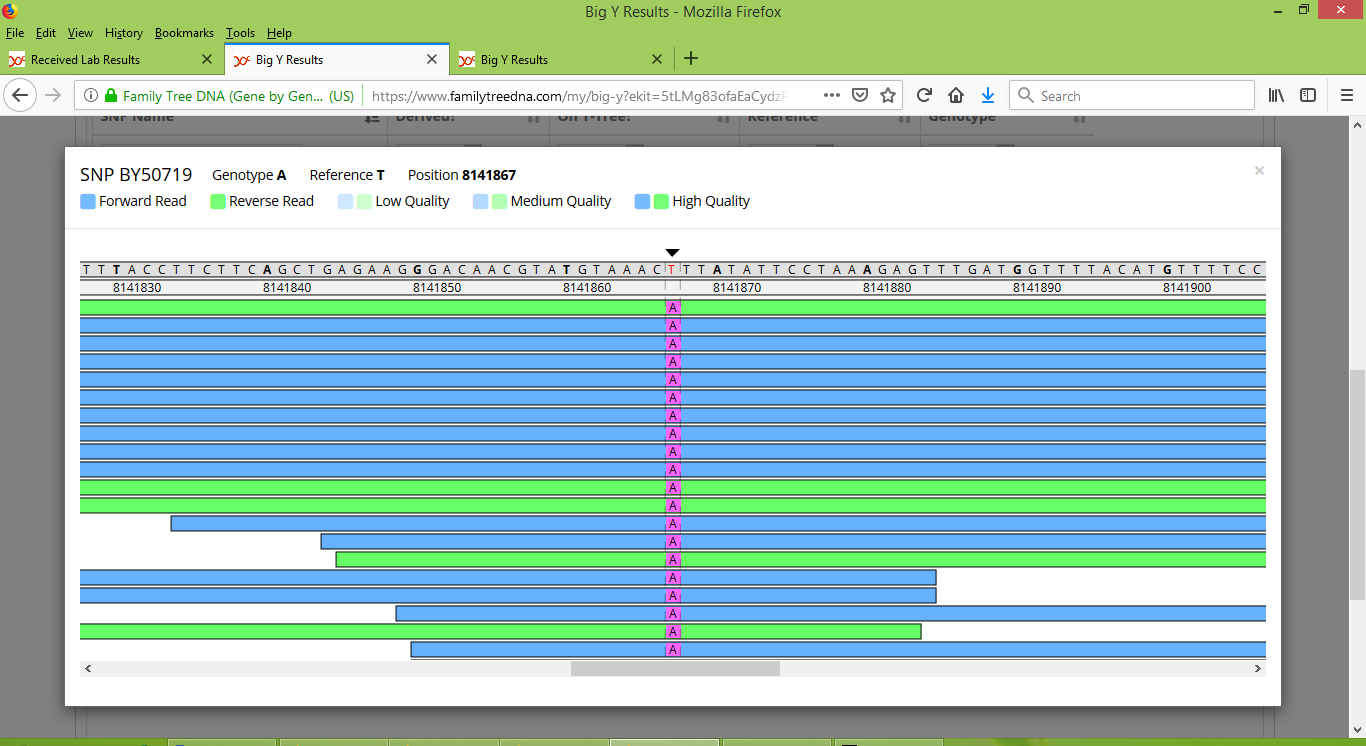
Thinking ahead, with future testing we may be able to determine which of Nathaniel or Samuel was the first to carry BY50719. All that it would take would be a DNA test performed on a man whose patriline traces back to one of Nathaniel's other sons. The presence of BY50719 in such a person would mean that Nathaniel is the BY50719 founder, whereas its absence would mean that his son Samuel is the BY50719 founder. In either case we can now be confident that Karen descends from Nathaniel, and so I have moved the display of her brother's STR results from the collection of "Matches that have no documented connection to James" to the collection of "Descendants of James' son James", as shown in the table of STR results for Group 6.
Here now is the current SNP tree for our project's Group 6, in which we are also able to show the position of settler James' son James junior. It is possible that he (i.e., James junior) is the founder for some of the 14 SNPs that are listed in the box just above him, although likely most of them arose in earlier generations.
Note that Russell, Gordon and Stuart each have a number of unnamed variants reported for them, for which the corresponding locations on the Y chromosome are shown in blue in the tree presented above. At such time as when these are found to be shared with other people, they will be assigned SNP names.
I mentioned above (when discussing Group 1) that analysing Big Y test results can be tricky. The results for Group 6 are especially so, particularly since there is an abundance of ambiguous and sometimes contradictory results that required manual investigation. In total 28 SNPs that Family Tree DNA had reported as being potentially present among some of Russell, Gordon, Karen and Stuart were inspected, assessed, and then set aside as not currently being informative for our Group 6. To convey an understanding of what I mean here, let's walk through an example, namely the SNP named BY26157.
Family Tree DNA reported that Russell is positive for BY26157 and Karen's brother is negative for it. Each of Gordon and Stuart were reported as a "no call". When we look at the reads taken for BY26157, for Russell we see a consistent collection of "A" alleles in its location (beneath the black arrow in the middle of the image below).
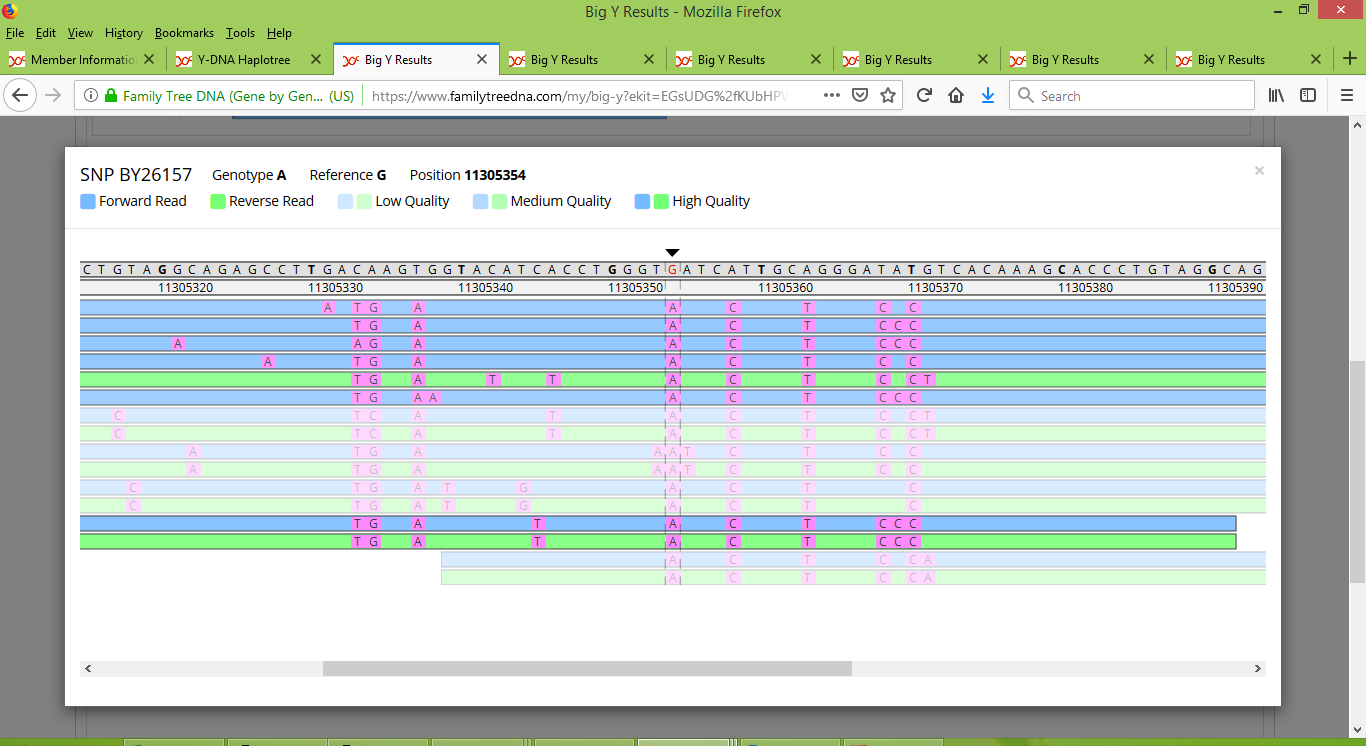
On the other hand, Karen's brother's results consist of just five reads, one that is positive and four negative:
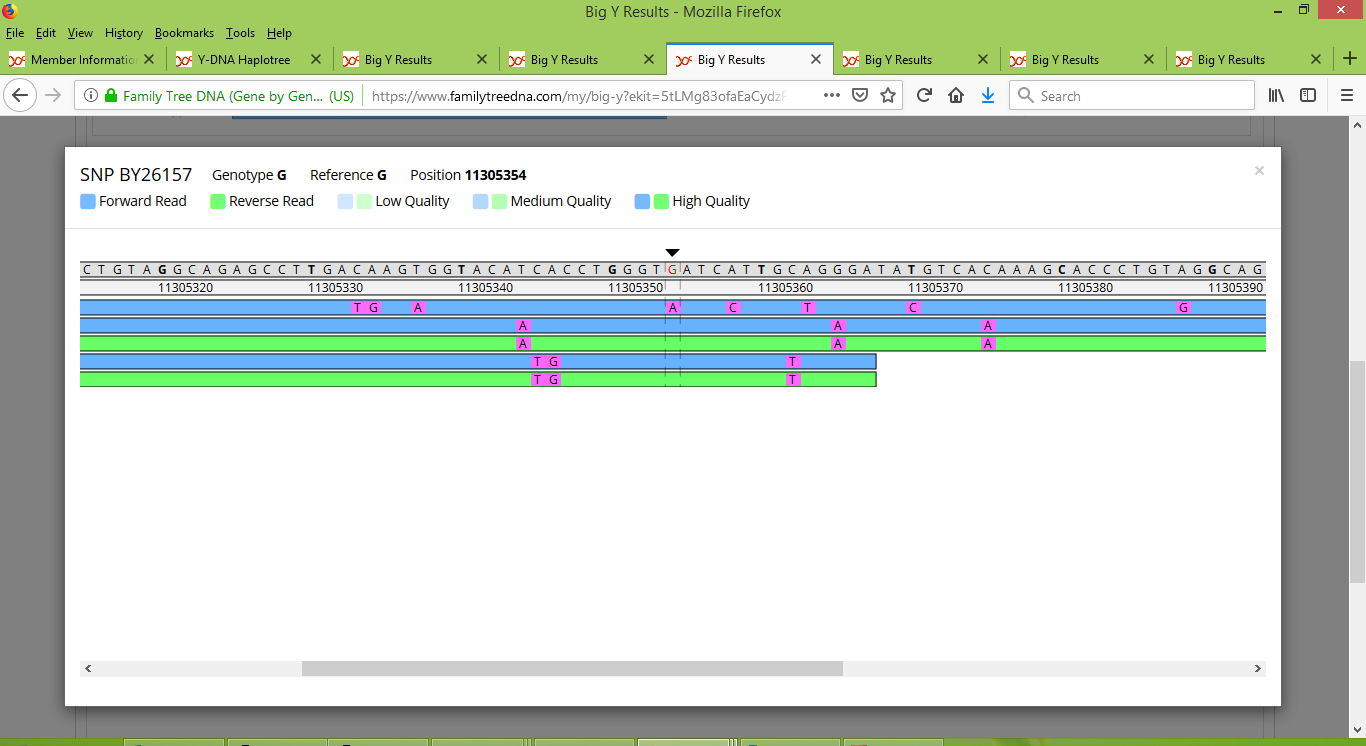
Meanwhile Stuart has a collection of low-quality reads that faintly appear to suggest that he is positive:
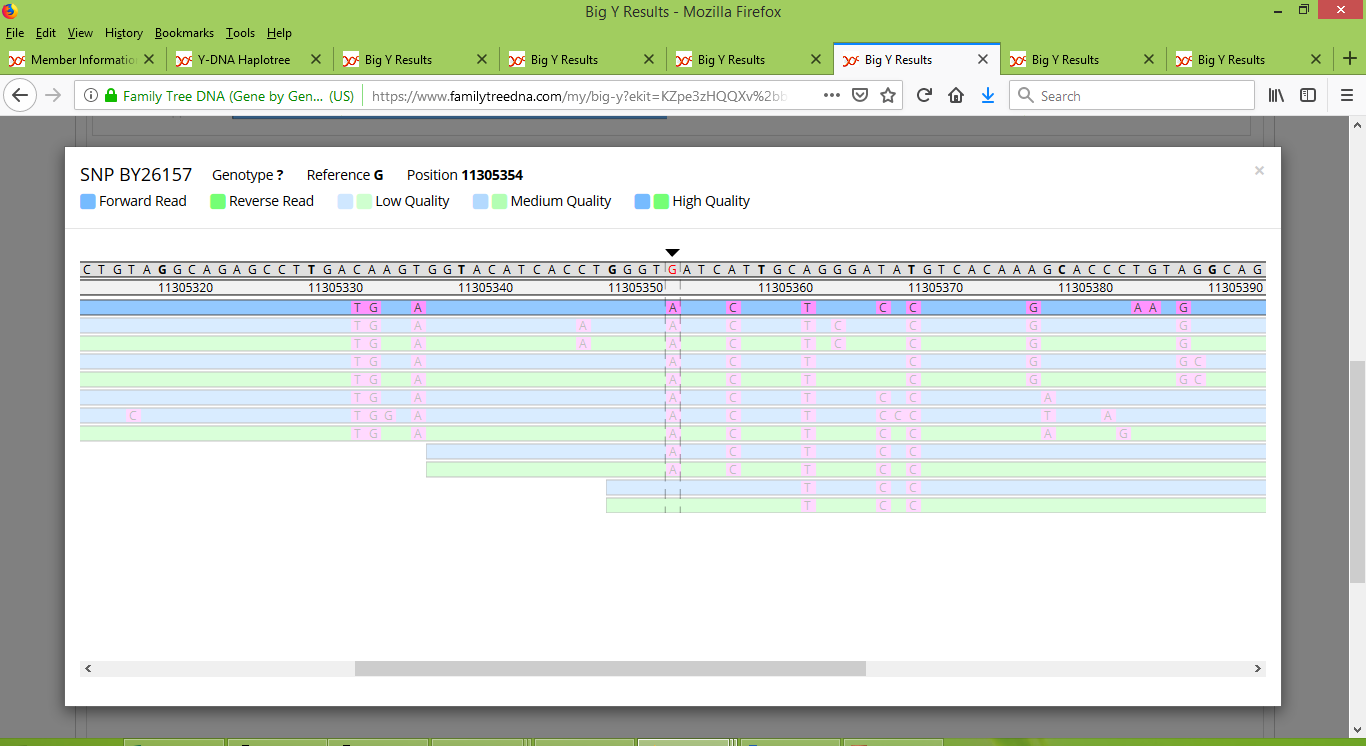
And Gordon has a mere two reads, both of low quality, faintly suggesting that he is positive:
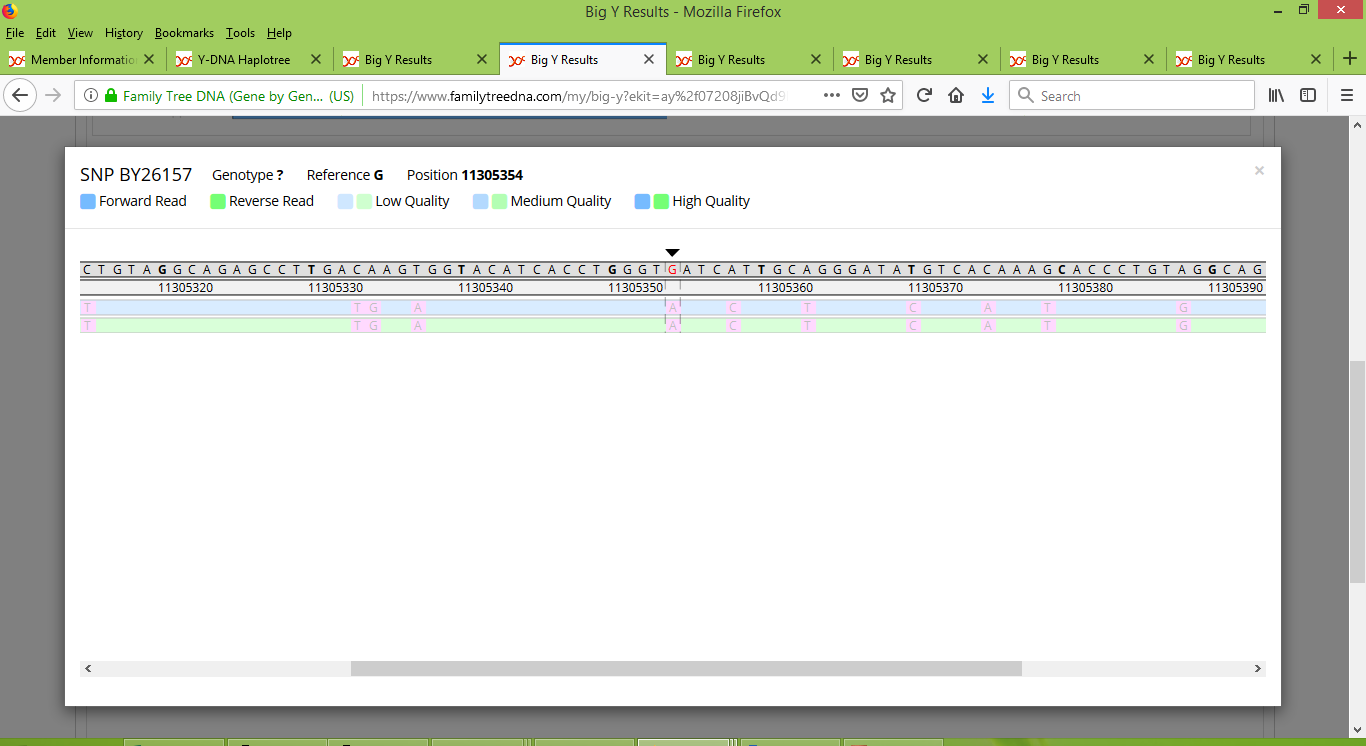
Many of the 28 SNPs that I've alluded to were like this, in that several of them involved "no calls", several of them had low quality reads, and several of them had very few reads. Moreover, their results often seemed to be inconsistent and contradictory. Continuing with BY26157 as a working example, positive results for Russell, Gordon and Stuart would indicate that their ancestor James junior would have carried the SNP. Having determined that Karen also descends from James junior (and in particular, from his son Nathaniel), then a negative result for her brother would suggest either an extremely rare backward mutation, or else a mis-call in genotyping. The mis-call seems to be the case at hand, given that only five reads were reported for Karen's brother, and one of the five suggested a positive result.
As for what testing would be good to do next within Group 6, as yet none of the descendants of settler James' son Jeremiah have done the Big Y test, so it would be interesting to start testing that branch of the family tree. It would also be useful to test some of the members of the group who do not appear to descend from settler James at all, but rather from some cousin(s) of his.
Groups 7, 8, 19
So far we only have one Big Y result in each of these groups, which is not enough to build SNP-based trees for them. Note, however, that in Group 8 there is a second Big Y test now in progress, which will allow us to begin to build a SNP-based tree in due course.Group 10
Our project's Group 10 primarily consists of people whose surname is McPike, McPeak, or something similar. Three of them now have Big Y test results, namely a nephew of Sandy (kit 18526), Taed (kit B4027), and an anonymous participant with kit number 49964. Sandy's patriline goes back to a Levi McPike who was born about 1780 in Pennsylvania. The identity of Taed's grandfather is uncertain, although he might have been a man named David Nelson who was born about 1906 in California (for more details, see Taed's pedigree details). Member 49964 has provided limited genealogy details, stating that he descends from a William Peak who was born about 1825 in County Down, Northern Ireland.Based on the three Big Y test results now on hand, here is the SNP-based tree for Group 10:
Sandy and Taed share a SNP named BY165777, which distinguishes their branch of the family tree from that of kit 49964. Meanwhile, all three share a collection of three SNPs (BY24089, BY24839 and BY24680) that currently appear to be characteristic of our project's Group 10 cluster. With limited genealogical details, we cannot yet indicate where in the tree specific ancestors should be placed.
All three of Sandy, Taed and member 49964 have some additional variants that are so far only identified by their locations on the Y chromosome (shown in blue in the tree above). As yet it is difficult to inspect these unnamed variants, although as more people get tested and these variants acquire names it will be possible to inspect them and determine their significance.
Group 20
Within Group 20, there are two people with Big Y test rseults. These were on hand at the time of the March 2018 update so there's nothing new to report for this group at the present time.Group 2
I've saved Group 2 for last, in part because it is the group for which we have the most recent Big Y test results. In total there are now six results on hand from members of Group 2. In March we had results for me (kit 23996), my father Angus (N21510), and Rodney (399984). Big Y test results are now also available from Tom (87744), Bob (454806), and kit IN25086 (who is anonymous).In the March 2018 update it was reported that Rodney had two unnamed variants at locations 14872255 and 15953286. These have now been assigned the names BY111567 and Y140924, respectively. Shortly after Family Tree DNA reported the completion of the three latest Big Y tests in Group 2, they were listing six other unnamed variants that had not been listed for Rodney back in March. I'm not sure just when these appeared in Rodney's results, but I'm happy to see that new variants may occasionally be found within people's results, even well after their test results were initially reported.
As it happens, several of these new unnamed variants that were listed for Rodney were also listed for one or more of the three new Big Y results for Tom, Bob and IN25086. It is because they were detected in the new results that I suspect FTDNA was then able to go back and re-assess Rodney's status at these locations. I contacted FTDNA about a few of these locations and shortly afterwards they were given SNP names. Moreover, some of these newly found SNPs are now reported as being carried by me and my father Angus too, even though the corresponding unnamed variant locations had not previously been reported for us. With SNP names now assigned to them, it is possible to use FTDNA's Y-Chromosome Browser to inspect them, and in so doing I believe that there is a straighforward explanation. In short it has to do with the number of reads at the locations, and the resulting levels of confidence about whether a variant is truly present. As an example, let's look at what began as an unnamed variant that was reported only for Rodney (i.e., it was not reported for any of me, Angus, Tom, Bob or IN25086) at location 16593778. Now that it has been assigned the name BY123278, it is easily inspected via the Y-Chromosome Browser, which shows that my test results had only a handful of reads at this location:
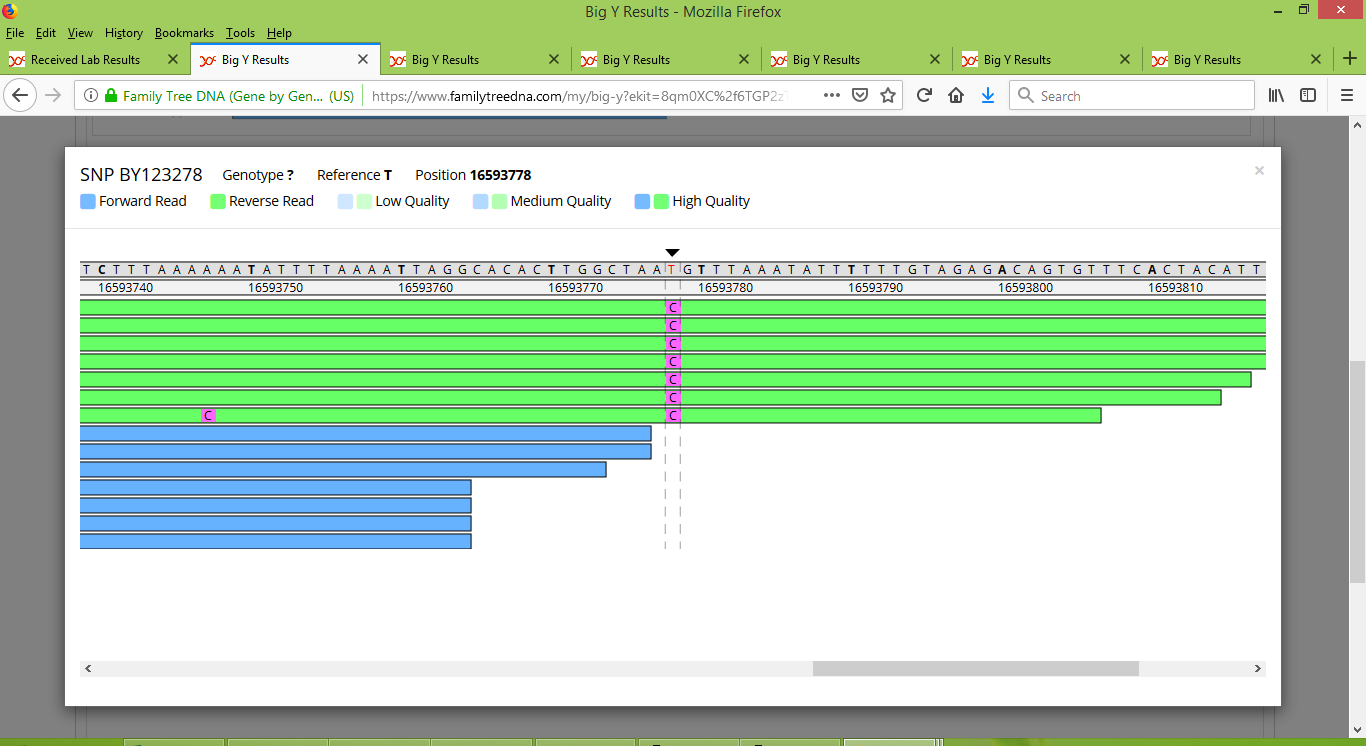
In contrast, Rodney's test results have an abundance of high quality reads that clearly indicate the presence of a variant:
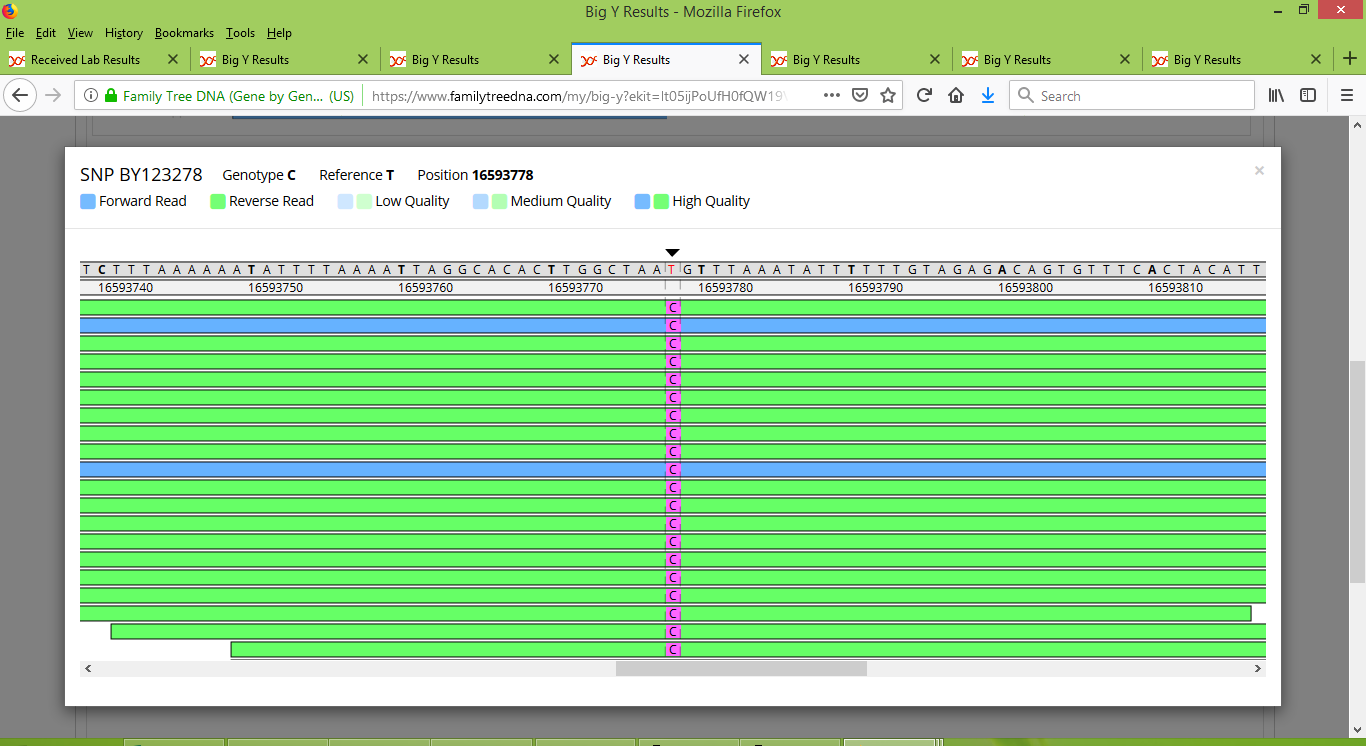
This situation is repeated for several other variants, which appear to have fallen below a confidence threshold to be reported when only one or two test results were considered, but a strong signal from other test results then pushed them into being noticed, re-assessed and duly named.
And it wasn't just with Rodney that this happened. Each of Tom, Bob, and IN25086 had at least one variant that was initially only reported for their own test results, but then got promoted to named SNPs that are shared by some or all six of the members of Group 2 to have done the Big Y test. As such an example, a variant at location 19039164 was initially reported just for Bob. It has since been given the name BY195776, and is a SNP that appears to be widespread within Group 2. It is easy to see how it could be overlooked among the other results though. For instance, my test results only contained three reads for this SNP's location:
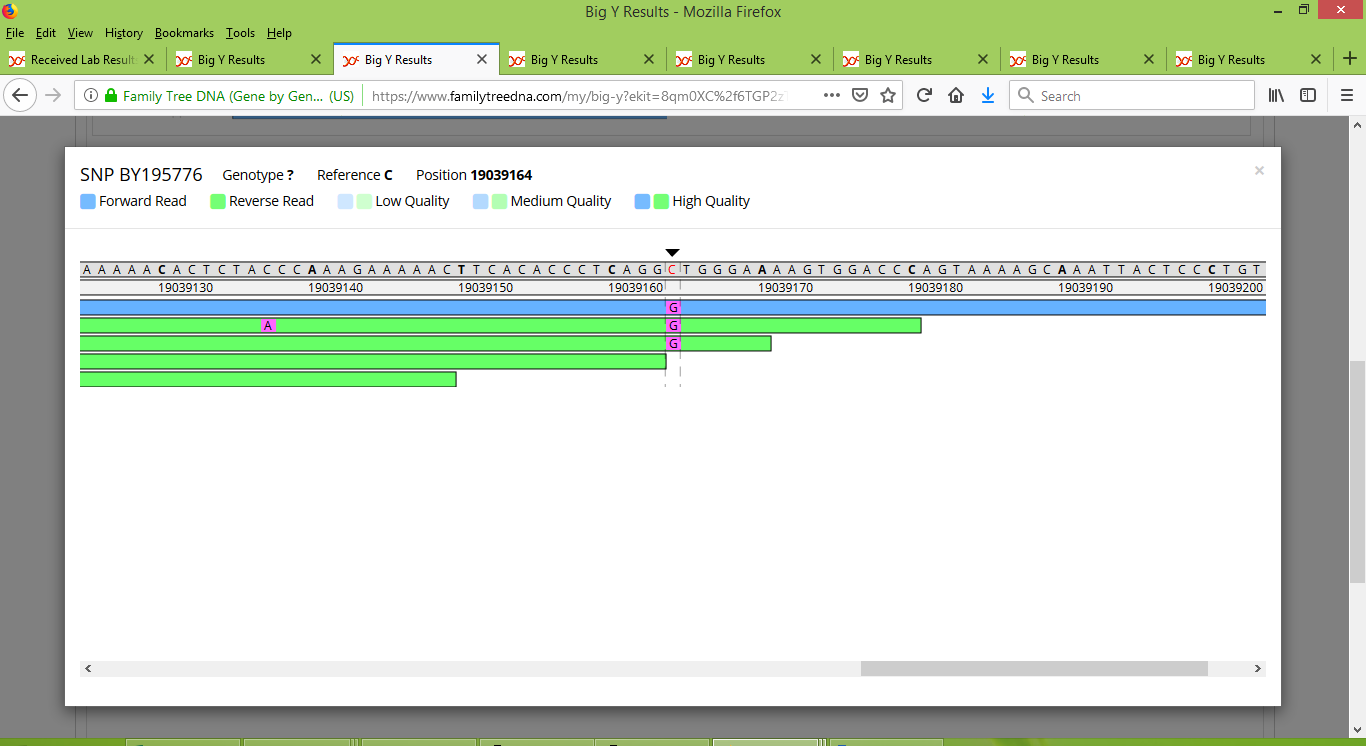
Even more extreme, Rodney has no coverage of this location. His test results have two reads that cover nearby locations, but these two reads both stop just short of covering the location for BY195776 (notice the black arrow just above its location).
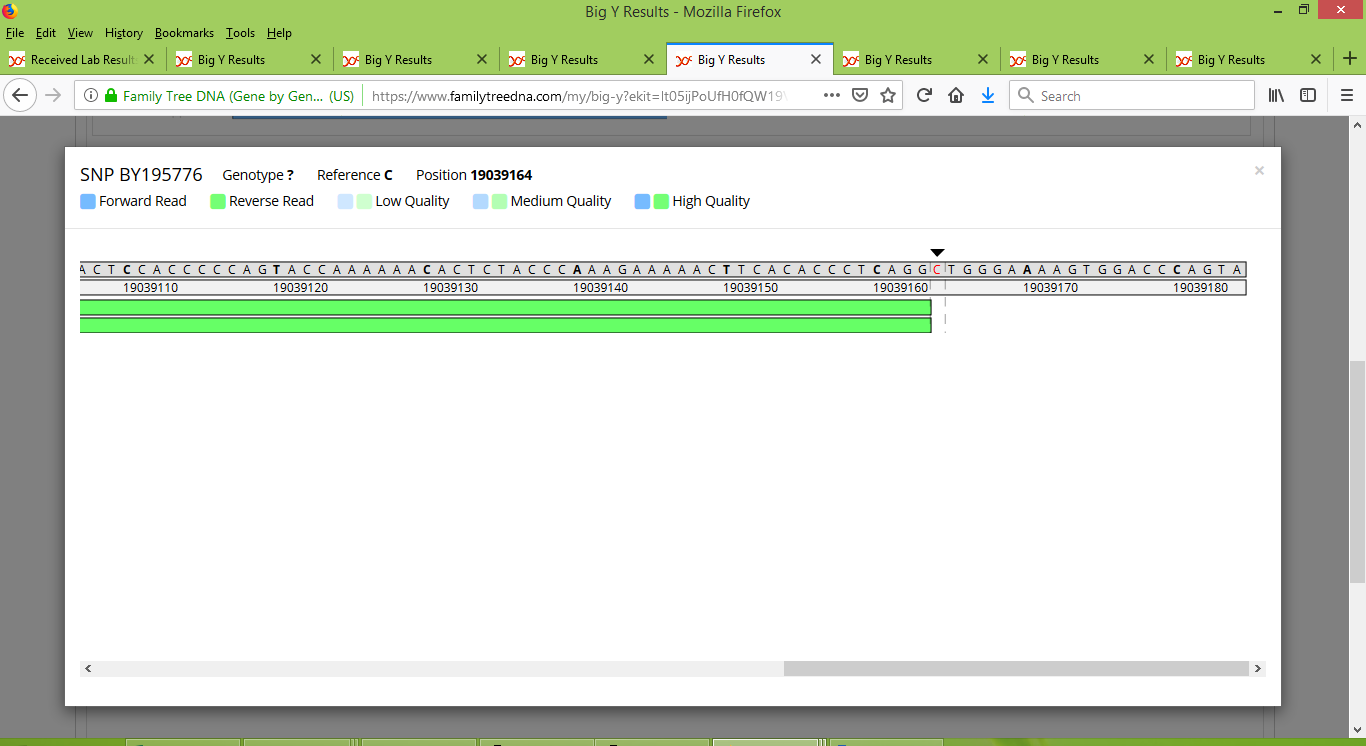
Only in Bob's test results were there enough reads at this location for it to stand out enough to get noticed. And we can see that they do indeed stand out in his results:
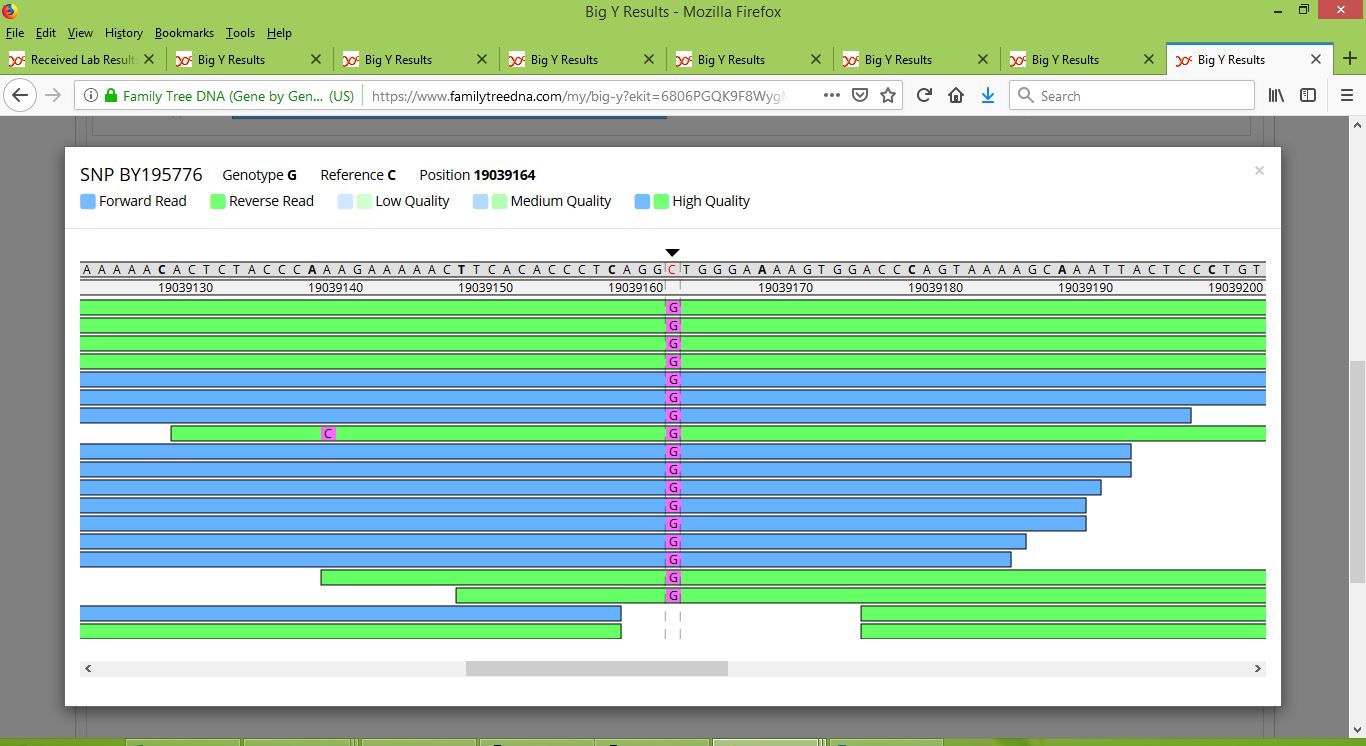
Something that I was hoping would happen, did happen, namely that we discovered a few new SNPs that are peculiar to only a portion of Group 2. For instance, the SNP now named BY111567 and found at location 14872255 is one for which only Rodney and IN25086 are positive, indicating that they belong together on a branch of the Pike family tree. To see what it looks like for a SNP to be positive for some people but negative for others, here is how BY111567 looks for Rodney:
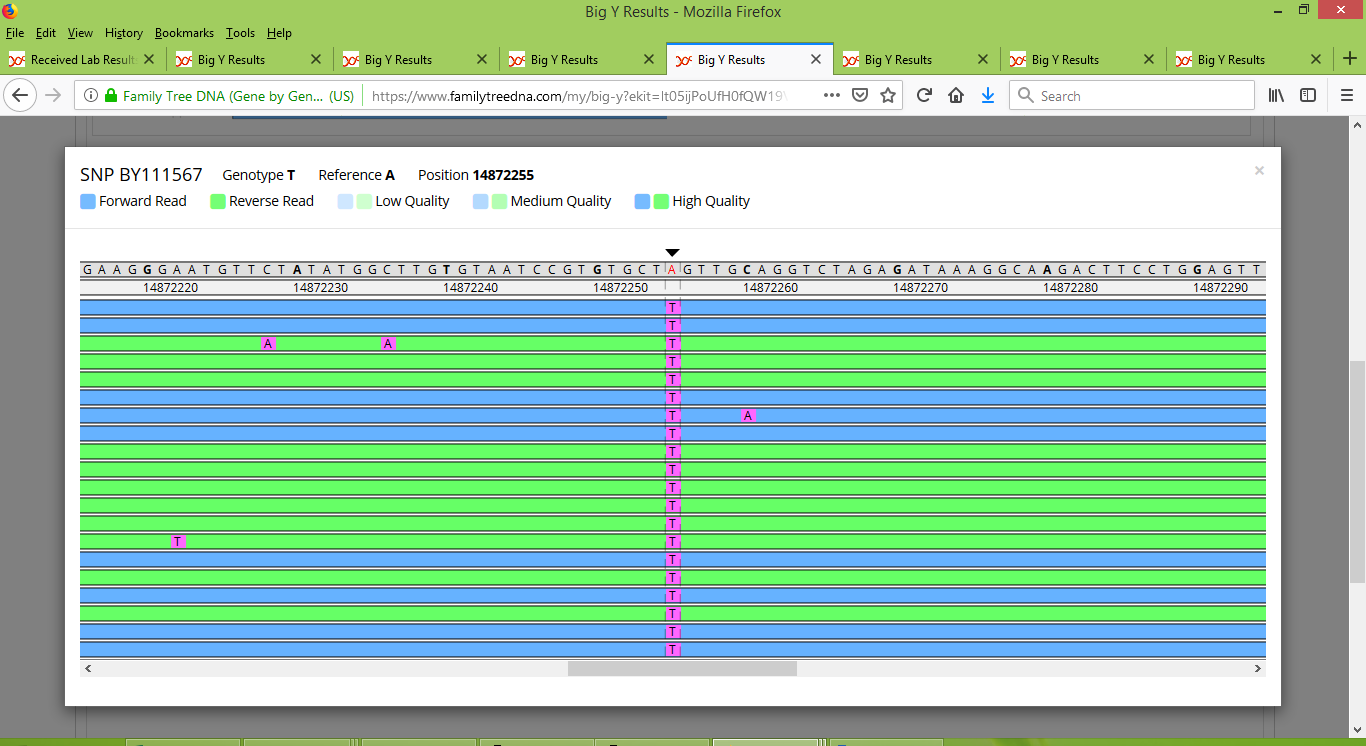
And in contrast, here is what BY111567 looks like for Bob:
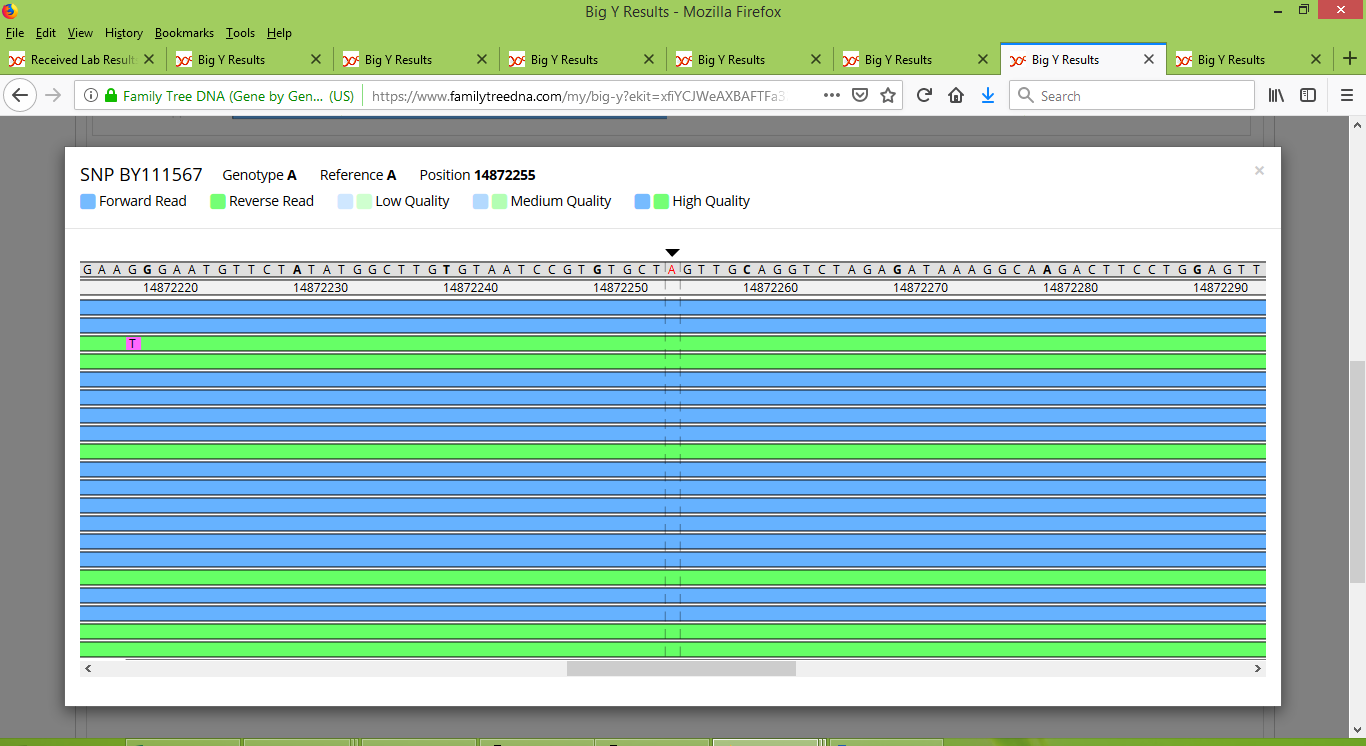
The SNP tree shown below illustrates the structure of the Group 2 family tree, as reconstructed from Big Y test results. As mentioned above, Rodney and member IN25806 fit together into a branch that is distinct from those that lead to Tom, Bob, Angus and myself, although we all have a common ancestor who carried the collection of 17 SNPs shown in the middle of the figure.
Who our most recent common ancestor was and when he lived remain a matter of inquiry, although as I've mentioned on previous occasions the general belief is that the Pikes of eastern Newfoundland descend from a mariner named Thomas Pike who married at Poole in 1680. A document from 1681 [reference number CO 1/47 (52i) 113-221 at the National Archives at Kew] shows Thomas present as a planter in Carbonear in Newfoundland, which is to say that he was overseeing several people engaged in the fishery (in fact, the document shows that he had 2 boats and 10 servants working for him). Despite working in Newfoundland, Thomas resided at Poole, as was typical of those involved in Newfoundland's migratory fishery. Thomas wrote a will on 20 March 1703 (click here to read a transcript of it) naming two of his four sons (i.e., Thomas and John). Within the following twelve months he died, as evidenced by an apprenticeship record from Portmsouth for his son William on New Year's Day 25 March 1705.
In the Big Y analysis done in March, the number of differences that Rodney had when compared to me and my father appeared to be substantial enough to suggest that the origin of our Pike family might pre-date mariner Thomas from Poole. There's a similarly large number of differences when me and my father are compared with the new results from Tom, Bob and IN25086. Most of these differences are with SNPs carried by me and my father, as if we have more mutations in our line than there are in other lines. Having now observed several more BigY results, what I'm now thinking is the case at hand is simply that FTDNA has been better able to identify new variants for me and my father because FTDNA has the benefit of two people's samples with matching variants. Moreover, results from father and son would be prone to discover more shared variants than when testing two men who are somewhere on the order of 5th to 10th cousins. And so it ends up appearing as though me and my father have more mutations than other people, not necessarily because we actually do, but because FTDNA has been better able to identify the ones that me and my father happen to carry.
Given this realisation, and also given that Group 2 now has three more Big Y test results on hand, I no longer think that our Big Y evidence is suggesting that our most recent common Pike ancestor lived substantially prior to mariner Thomas from Poole. Although me and my father still have several SNP differences when compared to Rodney, we have a similar number of differences when compared to Tom, and only slightly more when compared to IN25086. Bob has several unnamed variants, but in terms of those that have been given SNP names, he too has a similar number of differences when compared to me and my father Angus.
Moreover, when considering named SNPs, none of Rodney, IN25086, Tom or Bob stand out from one another as being unusually distinct (and hence from a distant branch in the family tree) from one another.
This is actually quite a relief for me, since the distant common ancestor theory that appeared to be suggested by the analysis done in March would, had it been true, have had a number of implications that would have been difficult to reconcile, such as why it is that a family with much older origins than mariner Thomas from the 1600s remains without genetic matches with other branches that do not have ties with Poole and Carbonear.
Now that we're back to the original scenario of having a common Pike ancestor who likely lived around the 1600s, there is still something curious about our DNA results that warrants some closer attention. If you look at the table of 111 STR marker results for Group 2, Rodney's results stand out because he has a number of STR mutations that he does not share with other members of the group who have tested those markers (such as marker numbers 9, 13, 16, 32, 60 and 72). This distinctiveness among Rodney's STR results had suggested that his branch of the Pike family was somehow different from the rest of Group 2, as if most of us are much more closely related to each other than we are to Rodney.
I mentioned at the beginning of this document that the Big Y test also includes some 500 additional STR results beyond the usual 111 STR results. There aren't yet many administrative tools to enable comparing these additional STR results, but I've been able to determine the full set of STRs that harbour differences and have gathered them into the table below. The first twelve of the 17 STRs are captured within the normal 111 STRs, so it is only the final five STRs below that come from the additional STR results of the Big Y test. One thing to notice here is that since only five additional STRs bear differences, then the vast majority of the the additional STR results are actually identical for all six men (when the STRs are inspected closely, 487 STRs show that all six have the same value, while another 48 do not reveal any differences but have some "no calls"). So even though Rodney seems to have an unusually high share of STR mutations within the group, there is nevertheless strong overall correlation between the sets of STRs.
| DYS389I | DYS389II | DYS458 | DYS454 | DYS576 | DYS442 | DYF406S1 | DYS413 | DYS481 | DYS446 | DYS540 | DYS650 | DYS389B | DYS508 | DYS512 | DYS514 | FTY1016 | |
| Rodney | 15 | 31 | 16 | 12 | 19 | 13 | 10 | 22-23 | 23 | 12 | 11 | 20 | 12 | 11 | 12 | 15 | 10 |
| IN25086 | 14 | 30 | 17 | 11 | 18 | 13 | 11 | 22-23 | 23 | 13 | 12 | 20 | 11 | 11 | 13 | 15 | 11 |
| Tom | 14 | 30 | 17 | 11 | 18 | 13 | 10 | 22-23 | 22 | 13 | 12 | 20 | 11 | 11 | 12 | 15 | 10 |
| Bob | 14 | 30 | 17 | 11 | 18 | 12 | 10 | 22-23 | 23 | 13 | 12 | 19 | 11 | 11 | 13 | 16 | 10 |
| Angus | 14 | 30 | 17 | 11 | 18 | 13 | 10 | 23-23 | 22 | 13 | 12 | 20 | 11 | 12 | 13 | 15 | 10 |
| David | 14 | 30 | 17 | 11 | 18 | 13 | 10 | 23-23 | 22 | 13 | 12 | 20 | 11 | 12 | 13 | 15 | 10 |
Regarding the colour coding in the above table, similar to the tables shown on our project's STR-based Results Page, the marker values with the yellow background colour are those that are nearly universal within the group. Shades of blue are used for values that drop below the universal ones, and shades of red for those that exceed the universal ones.
Although STRs are less reliable than SNPs because STRs are more prone to mutation (and sometimes these mutations are in parallel, or they can be the reverse of a previous mutation), it is nevertheless interesting to observe some things. For instance, the STR named DYS512 shows that Rodney and Tom share a value of 12, while the other four Big Y results show a value of 13. At first we might suspect that this indicates that Rodney and Tom belong together on a branch of the family tree. But then what about IN25096, who we now know from the SNP results to be closer to Rodney than Tom is to Rodney? The only sensible answer is that DYS512 must have mutated twice. Possibly once in Tom's line, and a second time along the portion of Rodney's line that comes after its split with IN25086, with both mutations taking a value of 13 down to 12. Or another possibility is that Tom, Rodney and IN25086 descend from a common ancestor who experienced a mutation from 13 to 12, and then later on another mutation from this 12 back to 13 happened within the portion of IN25086's line that comes after its split with Rodney. Right now it isn't easy to tell or guess which scenario is more likely.
However, DYS481 seems to be telling us something. And since it is the 58th of the usual STR markers we have the benefit of results from some additional members of our project, as seen in the table of 111 STR results for Group 2. Here we see that Rodney, IN25086, Bob, as well as Patrick (kit 504580) share a value of 23 whereas six other members of our project have been found to have a value of 22. From this it would appear that Rodney, IN25086, Bob and Patrick belong together within a branch of the family tree while the other six men belong to a separate branch. That is, assuming that this STR only experienced one mutation. If there are multiple mutations, then several other scenarios become possible. My hope is that more members of Group 2 will upgrade to test at least 67 STR markers (or, even better, upgrade to the Big Y test) so that we might be able to better understand this and other branching points in our family tree.
Getting back to genealogical matters for a moment, I want to mention something special about Tom's lineage. Of the many Pike lineages found at Carbonear in Newfoundland, his is (so far) the one and only one that I can successfully trace back to mariner Thomas of Poole on a generation by generation basis (incidentally, there is a line that I have been able to trace from mariner Thomas down to Pikes living in the English counties of Essex and Kent, but they're not yet involved with our project). Tom's 4x great grandfather Francis Pike died at Carbonear in 1835, where he is buried alongside his wife Jane and some of their children. You can view their gravestones by clicking on this thumbnail image:
Francis himself was born about 1760, although I don't yet know where. Through a combination of records I've been able to determine that Francis had siblings named William, Edward, Catherine and Mary. And while I still do not know the name of their father, I have been able to determine that their grandfather was John Pike who was baptised at Poole in 1687 to mariner Thomas and his wife Susanna Bird (they were married at Poole on January 17th 1680).
Where mariner Thomas was born is yet another mystery. I'll be making visit to the UK National Archives sometime in 2019 to look at some records that I hope will contain some clues in this regard.
Some Closing Comments
One of my opening remarks is worthy of being repeated: our DNA project is very much a team effort, and is only made possible by the participation of many Pikes from various branches of our family trees. We welcome new members to join our project, and we encourage existing members to upgrade their results to the extent that they are able to do so.
It is also interesting to have observed that investing in a Big Y test produces results that are not necessarily static. Although the underlying "reads" will remain unchanged, it may well be that some previously unreported mutations will be reported as time passes, particularly in cases in which the reads for a mutation did not surpass a threshold for being initially reported (perhaps because of too few reads) but are later found to be significant when detected in another person's results. This also underscores the group effort that is required when it comes to getting the most out of genealogical DNA testing.
Last Modified: Monday, 24 December 2018, 08:49:33 NST