This update concerning the Pike DNA Project is going to be long and it's going to be technical, but I hope that you will also find it interesting and informative. This message is going to be devoted entirely to test results from Family Tree DNA's "Big Y" test. This is not the usual 37 or 67 or 111 STR marker test for which we show results on this webpage.
Rather, it is a test that FTDNA introduced in late 2013 and looks at about ten million SNP markers on the Y-chromosome. SNP markers (i.e., Single Nucleotide Polymorphisms) are the type of genetic marker that are used to define Y-DNA Haplogroups (such as R1a, R1b, etc.) and their subgroups. And when we get into enough nested levels of sub-sub-sub-...-subgroups we are dealing with SNPs that are specific to particular surnames and even to particular branches of a family tree. It has been my hope for a long time that we would be able to make use of the power of the Big Y test within our project to help identify the structure of some of our family trees, even in cases where we don't have historical records to tell us the shape of the tree. We're now at a point where I believe we are beginning to reap this reward.
A number of our project members ordered Big Y tests during the sale that FTDNA had in late 2017. Their results are now all in, and I will discuss them and what they mean for the groups within our project. And by "groups" I mean the genetic clusters that we've numbered within our project as Pike Group 1, Group 2, etc. So far only four of our groups have more than a single Big Y result to work with, namely Groups 1, 6 and 20 (each with two Big Y results) and Group 2 (with three results). So it is these four groups that I will concentrate on in the following discussion.
In terms of what you will see on our public website regarding Big Y results, let me start by saying that a full list of each person's ten million marker values will NOT be put on display. Rather, what I aim to do is to develop SNP-based trees that show when each SNP-based mutation arose, and the SNPs that have subsequently accumulated within the different branches of each group's family tree. These trees will, out of necessity, be limited to those individuals who have taken the Big Y test, because it is only those individuals for which we have sufficient SNP results to work with.
Before I talk about any actual results though, I should describe how FTDNA reports them. Of the roughly ten million markers that are analysed with the Big Y test, it is those that are found to be "derived" that are of interest, namely those which differ from the original value that humanity carried long long ago. Derived markers are described in two ways: either as named variants or unnamed variants. Up until sometime last year FTDNA was referring to "unnamed variants" as "novel variants" but they have since revised their terminology. When somebody is found to have a derived value at a location at which nobody has previously been found to have a a derived value, then since the discovery is new the location is simply identified by its position on the Y-chromosome, such as position 12910866. In time, and especially if this variant is found in other men, it may be given a name. In the case of position 12910866, it has since been given the SNP name L554.
To further explore one's results, FTDNA provides a "Big Y Browser" tool that visually presents the raw data from the Big Y test. A screenshot of my results for L554 and the other positions in the vicinity of position 12910866 can be seen below. Each horizontal line in the figure corresponds to a separate scan of this portion of my Y-DNA, which (in this example) consists of scans that were deemed to be of high quality. Most important though, notice that I carry a G nucleotide at position 12910866, whereas the reference value is an A nucleotide.
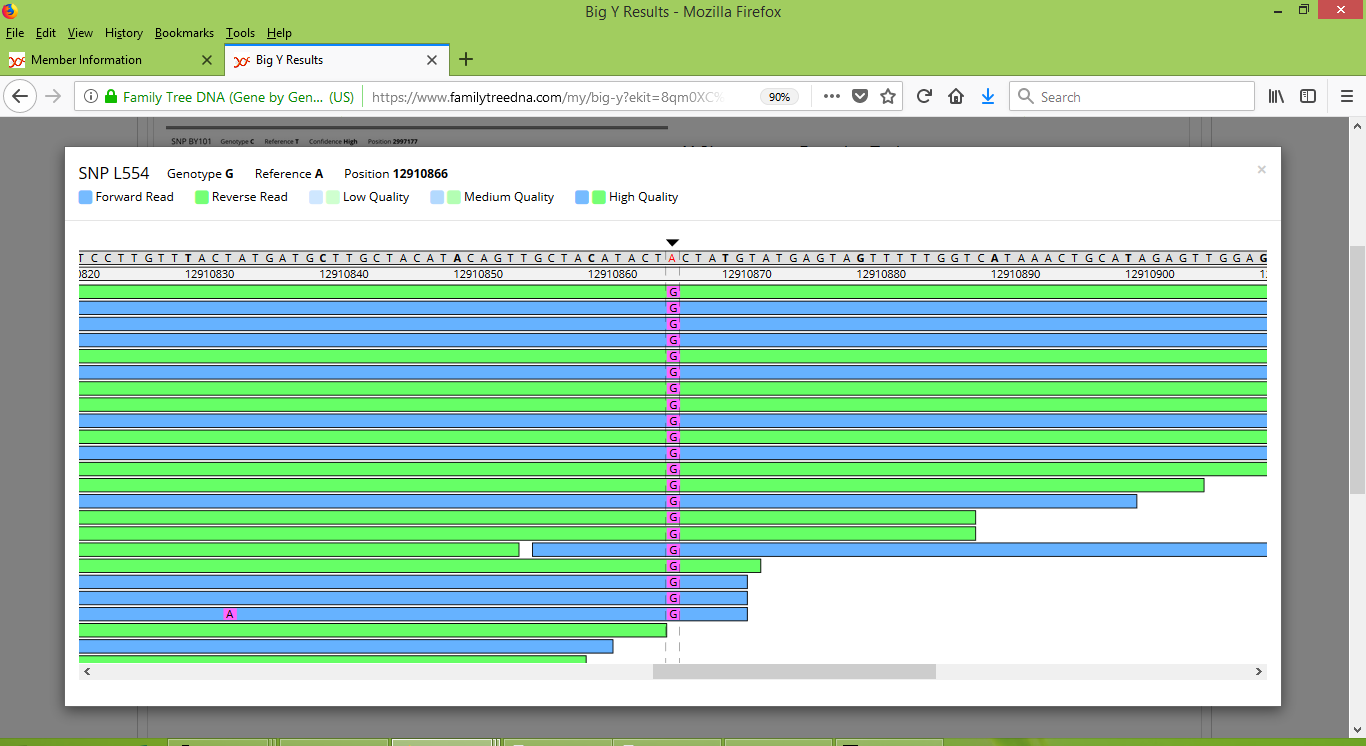
To continue using myself as an illustrative example, FTDNA currently reports that I carry 934 named variants and five unnamed variants. When my results were initially reported, I had a larger number of unnamed variants, but several of them have since been given names. This transition appears to have been semi-automatic, although my suspicion is that it was performed by staff at FTDNA as part of the normal procedures when new Big Y test results are produced. That is, when other people such as the two others in my Pike Group 2 were found to also carry some of my previously-unnamed variants, the technicians at FTDNA were able to identify the variants as phylogenetically meaningful and then proceeded to assign names to them. Having watched the Big Y results of several of our project members, this naming process appears to happen about four or so weeks after an unnamed variant is found to be carried by more than a single person. So really it is an ongoing process that is driven by the addition of new test results.
The real power of DNA testing is realised when comparing with other people. Currently FTDNA displays two Big Y matches for me, namely the two other men in our Group 2 who have also done the test. FTDNA shows the variants (both named and unnamed) at which we differ. For one of these two people, there are four variants that separate us, and for the other there are 14 variants. Usually each variant in question represents a location at which one of us has the ancestral value and the other has a derived value. It can sometimes take a bit of work to determine which person has the ancestral value and who is derived. The derived values are the exciting ones, as they represent new SNP mutations that would have occurred along the patriline of the derived person sometime after it branched away from the patriline of the non-derived person.
Another thing that I think is great about Big Y testing is that because SNPs are so much more stable than STRs, we can now calculate some much better estimates for when shared ancestors lived, namely TMCRA which stands for the Time to the Most Recent Common Ancestor. Yes, our usual 37 or 67 or 111 STRs can be used to calculate TMRCA estimates, but we now have another basis for doing so, and one that I think can be better calibrated. I'll talk more about calibration soon, but the general estimate that appears to be getting mentioned in places such as this blog post by Roberta Estes, is that an average of one Big Y SNP mutation will occur each century within any given patriline. So, for example, if two men are separated by 14 variants, then the average scenario would be that they each accumulated 7 distinct variants in the time since their most recent common ancestor, which in turn would suggest a TMRCA of about 700 years.
On the matter of calibration, and the general estimate that one mutation will occur per century within an average lineage, let's take our own look at what we can determine from the results within our project.
Group 1
Within Group 1, which is our project's largest group, so far two people have taken the Big Y test, namely Larry (kit number 129135) and Roger (kit 70909). Larry and Roger both have well-documented pedigrees that trace back to John Pike who was born in or near Whiteparish in Hampshire and who settled in Massachusetts in 1635. In particular, Larry descends from John's son John, and Roger descends from John's son Robert. Moreover, Larry and Roger are in the same generation, each being 9th great grandsons of John, who himself we think was born about 1572. As that was 446 years ago, the "one mutation per century per lineage" estimate would tell us that we would expect Larry and Roger to be separated by about nine Big Y variants. The reality is that FTDNA reports that they are separated by five variants. One of these five is named BY28714 while the other four are unnamed variants at positions 9000668, 13545985, 13547050 and 17359321. As shown in the following screenshot, when using the Big Y Browser to inspect each of their raw results for BY28714, Larry and Roger have a smattering of medium and lower quality results that make it difficult to conclude whether they carry the BY28714 variant.
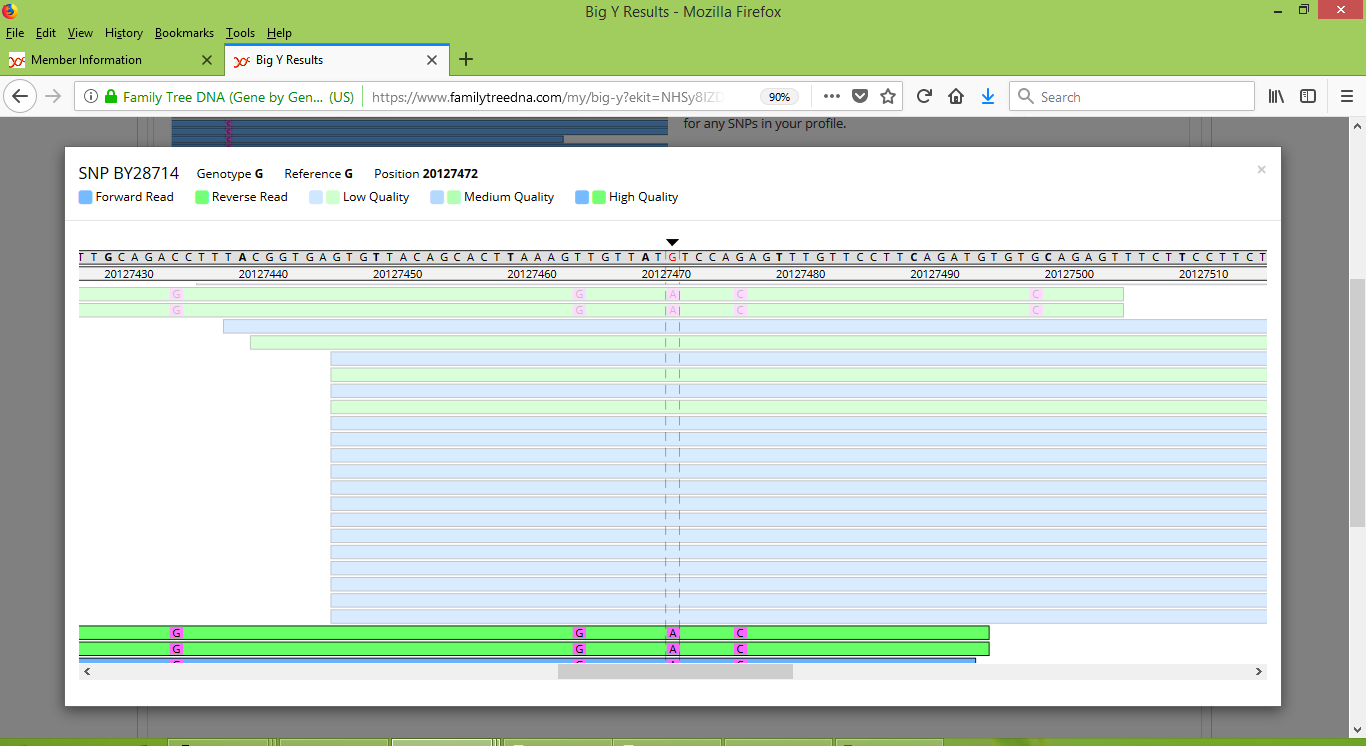
As it happens though, both Roger and a more distant non-Pike Big Y match of his are reported as carrying the BY28714 variant, which indicates that it is a SNP mutation that happened long ago, likely well before our Group 1 Pike family first acquired the Pike surname. So most likely everybody in our Group 1 has inherited the BY28714 variant, including Larry (even though his raw results look slightly ambiguous for this variant). It would therefore follow that Larry and Roger are only separated by a total of four observed variants. So based on the evidence that we have from our Group 1, it appears that instead of an average of one mutation per century per lineage, we are seeing an instance of an average of about 0.45 mutations per century per lineage.
Bear in mind that we are working with an extremely limited sample size here, namely just one pair of men, and it could well be that our one example is an extreme outlier when compared with the overall reality across all families. Only as more people engage with Big Y testing will we be able to improve the statistical significance of the results that are available at any given time.
I mentioned earlier that SNP-based trees were something that we could develop from Big Y test results. Below is the first such instance of a SNP-based tree for our Group 1. We have long known that our Group 1 fits into Y-DNA Haplogroup R1a, a large portion of which carries the SNP named M198. We have also known for a while now that Group 1 belongs to the sub-...-subgroup of M198 that is based on the SNP named L664. More recent information has shown that a collection of six SNPs (YP5461, YP5462, YP5463, YP5464, YP5465 and BY30049) represent an even more refined subgroup. Right now it is not known what order these six SNPs arose, although possibly that will be discovered in time. Based on the Big Y results of Larry and Roger and the non-Pike that is listed among their Big Y matches the SNP named BY28714 appears to be the next SNP to have arisen during the evolution that led to our Group 1 Pike cluster. Then comes the split with the line for this non-Pike; he is separated from Larry and Roger by 16 and 15 variants, respectively, which seems large enough to suggest that this branch in the tree pre-dates the advent of the Pike surname. And then there's the point at which Larry and Roger split apart, which we know happened with the birth of two sons of John Pike who settled in Massachusetts in 1635. As it happens, Larry has only one further variant, not yet named but at position 9000668. And Roger has three, also currently unnamed. We do not yet know the chronological order in which Roger's line accumulated these three variants, but as more members of the family get tested we may be able to pinpoint when each one arose.
As we make advances with pinpointing the origin of each variant, we may be able to identify which Pike man was the first to carry each of these variants. Each of his descendants ought to carry his variants, which also means that when somebody without a well-document pedigree does the Big Y test we will be able to rely on matching variants to determine where the newly tested person's line belongs as a branch in the family tree.
Something else worth noting is that Larry and Roger both have additional unnamed variants that are reported by FTDNA, but which are currently not being counted as differences between them. Honestly I'm not sure why this is the case, although I can speculate that it may be that some of these variants weren't able to be assessed for both of them, similar to the situation of a "No Call" that can happen when genotyping at a particular location fails. Again, as more people get tested, we may be able to determine the placement of these variants within the SNP tree. Their eventual inclusion may also result in some adjustment to the small calibration exercise that we went through for estimating TMRCAs.
Group 20
Let's now turn our attention to Group 20, which currently has only four Pikes with STR-based test results. All four of these men have ancestry that traces back to the vicinity of New Harbour in Trinity Bay, Newfoundland. Moreover, all four of them descend from men named George Pike. George was a popular forename in this family, which has made sorting out the different family branches a challenge. The loss of relevant church records has also been troublesome. My suspicion is that this Pike cluster hails from a George Pike/Pick who came from Christchurch in Hampshire and who is known to have married in Newfoundland in 1794.With regard to Big Y testing, we have results for Ernest (kit 247419) and Stephen (343554). They appear to descend from two different men, both named George Pike and both likely born about 1820. We can speculate that these may be two grandsons of the George Pike who married in 1794. If so, that would mean this elder George is the MRCA, and supposing he was born about 1770 then we have a TMRCA of about 248 years. If the estimated average of "one mutation per century per lineage" holds for this family, then we would anticipate that Ernest and Stephen would be separated by about five variants.
What we find is that FTDNA reports only two variants for which they differ. As with our Group 1 then, it seems that the evidence we have suggests that this estimation of 1.0 may be too high. To observe a total of two mutations among two lines over a period of 248 years would suggest an estimate of 0.40. I will nevertheless remind everybody that our sample size is far too small to be relied on as statistically significant.
Ernest and Stephen have a further one and five unnamed variants respectively, but which aren't reported as differences for them. I suspect these six variants may have the status of a "No Call" with the one of Ernest or Stephen, but at present the Big Y Browser tool doesn't enable us to inspect unnamed variants. As more people test and we learn more about these variants, they may come to further define the SNP tree for the Group 20 Pike family.
Neither Ernest nor Stephen have any other Big Y matches, so we do not currently have the benefit of a non-Pike to help us to determine which of Ernest and Stephen's shared variants arose before versus after the founding of the Pike surname for their family. Said another way, Ernest and Stephen share 23 named variants that cannot yet be chronologically ordered. Only when some men who share some (but not all) of them will these 23 variants begin to be placed in relative order to one another.
Something else to point out is that among these 23 variants, several of them have names that begin with the letters BY, which indicates that it was Family Tree DNA who named them, and that doing so was made possible because they were discovered through Big Y test results. In addition to the insights that we obtain regarding our Pike family origins though DNA testing, our Big Y test results are also contributing to the bigger project of discovering and mapping out the many branches of the human Y-DNA SNP tree.
Group 6
The next group that I want to consider is Group 6, which consists of descendants of James Pike who arrived at Charlestown in Massachusetts sometime in or before 1647. It is not now known where he came from.The two men in this group with Big Y results are a brother of Karen (kit number 61277) and our project's co-administrator Stuart (48191). Stuart has been able to document his ancestry all the way back to settler James. However, Karen has gotten stuck in tracing her lineage and can reliably only get back to a George S. Pike who was born in 1823 at Glens Falls in Warren County, New York. There is strong suspicion that George is closely related to a Samuel Pike who was born about 1730 at Holliston in Massachusetts and who resided in Warren County from about 1812 up until the time of his death in 1838. Karen has successfully traced Samuel's ancestry back to settler James. But alas, no records to confirm how George S Pike is related to Samuel Pike have yet been found. More genealogical details about Karen's research into her ancestors can be found on this webpage.
If we surmise that George's father was a son of Samuel, then Karen and her brother would be 7th cousins to Stuart's father. Their Pike lines would have diverged when settler James' son James Jr had two sons: Nathaniel in 1685 (leading to Karen) and Samuel in 1690 (leading to Stuart). With James Jr being born in 1647, that yields a TMRCA of 371 years. Referring again to the estimated average of "one mutation per century per lineage" we would therefore anticipate about seven Big Y variants to separate Stuart and Karen's brother. But if our calibrations from Group 1 and 20 are more typical, where we've witnessed estimates of 0.40 and 0.45 mutations per century per lineage, then with a TMRCA of 371 years we would expect something on the order of three Big Y variants as differences in the test results for Stuart and Karen's brother.
What FTDNA reports, however, is nine variants, which exceeds all of these estimates. This would, at first glance, appear to suggest that Karen and Stuart are more distantly related than would be the case if George S Pike were a grandson of Samuel Pike who lived 1730-1838. Indeed, it might even suggest that the most recent common ancestor for Karen and Stuart was not settler James but some ancestor of his. That would be exciting in its own right, for it would indicate a second immigrant to the USA from this Pike family, possibly providing a second opportunity for us to investigate and discover where the family originated prior to its arrival in the USA.
As it happens, four of the nine variants that are indicated as mismatches between Karen's brother and Stuart are variants that have already been given names. That enables us to inspect them more closely with the Big Y Browser tool. One of these variants is BY26256, and when we look at Stuart's raw data for it, here is what we see:
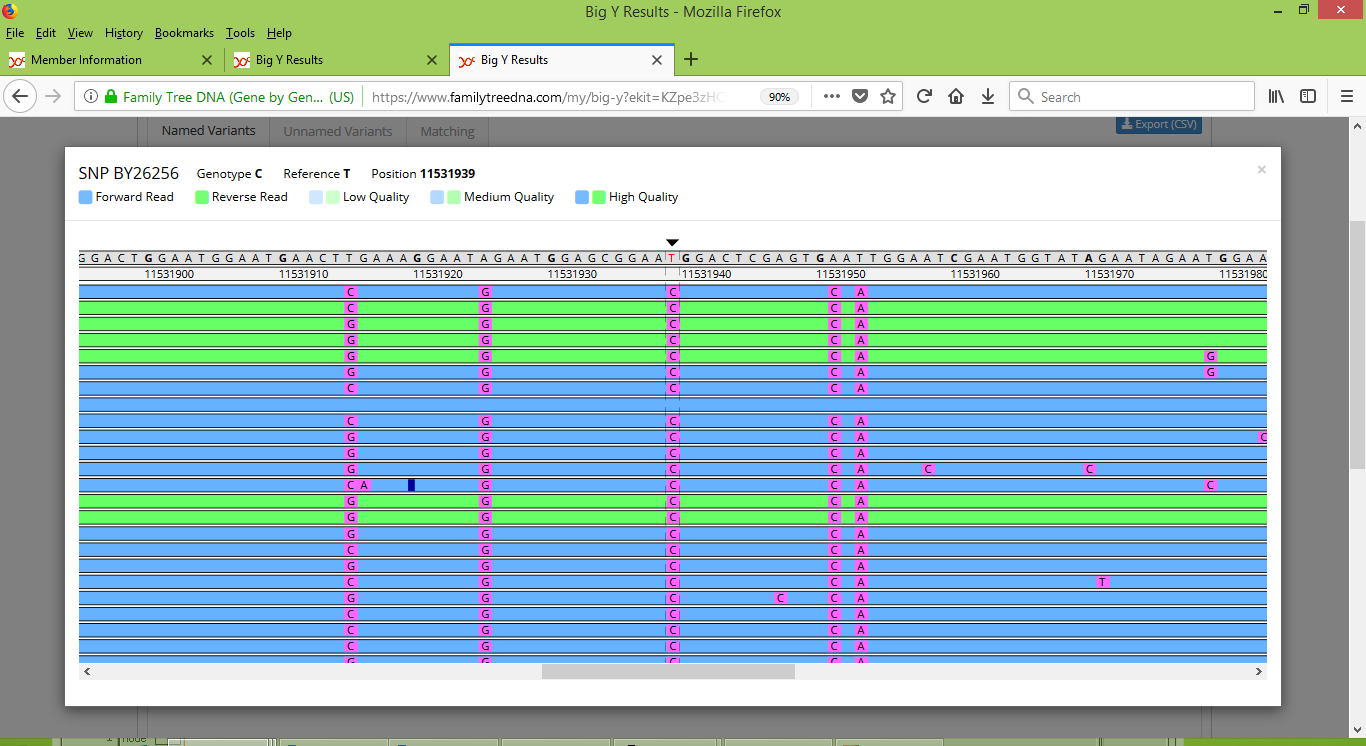
And here is the correspond view for BY26256 for Karen's brother:
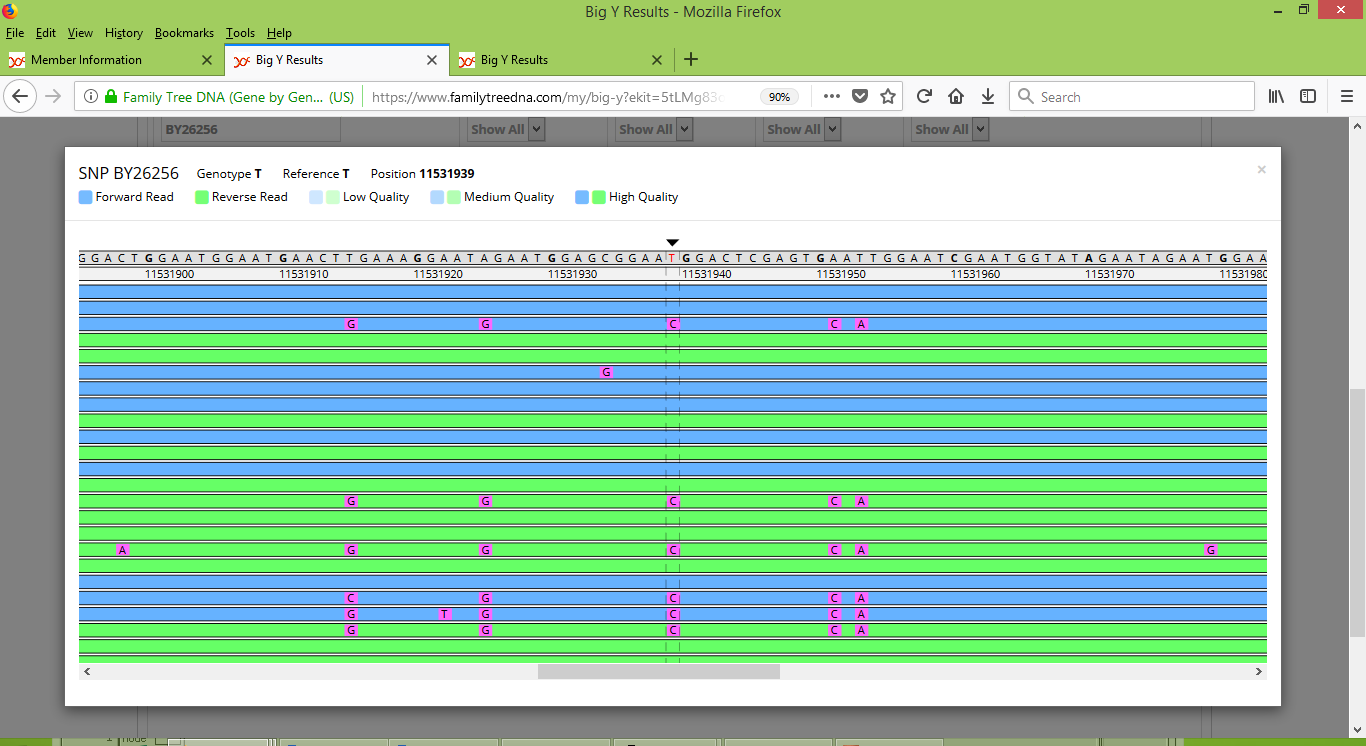
Observe that for Stuart, there is strong and consistent detection of a mutation at the position for BY26256, as well as for four nearby locations as well. For Karen's brother these mutations are sporadically and inconsistently reported among the several scans that covered this region of the Y chromosome. By doing some searching at the ISOGG YBrowse website I was able to find that some of these nearby locations of interest have also already been given names. In particular, locations 11531925, 11531951 and 11531953 have been respectively named BY26255, BY26257 and BY26258. Armed with SNP names, it is straightforward to query and discover that FTDNA has assessed two of these three SNPs as No Calls for Karen's brother and has reported the third as Negative (i.e., not derived). For Stuart, one has been reported as Positive (meaning derived) and two as No Calls (which seemed odd, since Stuart's results appear to be strong and reliable).
I contacted FTDNA about what looked to be inconsistency in how these results were being assessed. It was agreed that there was some inconsistency happening in this case, possibly due to some misalignment in part of the analysis procedure. I was further told that FTDNA would look into how to better filter out situations like this.
That's an answer that I'm happy with. This is a relatively new technology that we are making use of, and all of us (including Family Tree DNA) are learning how to work with it and get the best information that we can from it. So I'm happy that we've been able to contribute towards refining the process and making this better and more informative for everybody.
And in the meantime, I'm also happy because this answer from FTDNA suggests that the tally of nine mismatching variants between Stuart and Karen's brother is likely to be reduced once FTDNA is able to implement the filtering that they said they would explore. Even though the nine mismatching variants don't include BY26255, BY26257 or BY26258 that were just mentioned as being inconsistent, I did spot some similar inconsistency among the nine. But BY26255, BY26257 and BY26258 made for the best example to convey in this report.
Given that the tally of nine differences in this case is prone to be adjusted downwards, then our initial impression about how distantly related Karen and Stuart appeared to be can be correspondingly adjusted. Instead of a TMRCA that pre-dated settler James, a more recent TMRCA now seems viable, and is in keeping with the theory that George S Pike is a grandson of Samuel Pike, even though we still do not have irrefutable evidence to this effect.
When the Big Y results for Stuart and Karen's brother were initially reported, Stuart was shown to have 28 unnamed variants and Karen's brother 27 (several of which matched those carried by Stuart). A few weeks later many of these unnamed variants had been assigned names. Today FTDNA only reports five unnamed variants for Stuart and six for Karen's brother. Among the variants that got named were those now called BY36048, BY36049, BY36050, ..., BY36060. Although neither Stuart nor Karen's brother have any other Big Y matches reported by FTDNA (which only reports people as matches if they differ by about 40 or fewer variants), within the Haplogroup R1b-U106 Project it can be seen that a fellow named Schupp shares these newly named variants. Thanks to Schupp's participation in Big Y testing, we can therefore conclude that these particular variants pre-date the origin of the Pike name for our Group 6 cluster. And thanks to Stuart and Karen's brother, a new sub-sub-...-subgroup based on these variants has been identified, with them and Schupp now being placed into it (without Stuart and Karen's brother, these variants would have remained detected within Schupp's results but unnamed). This is great example that nicely showcases how our individual contributions to this new science are working towards the greater good.
Presented below is the SNP tree for Group 6. It shows the collection of variants shared with the non-Pike Schupp, as well as the collection of variants that arose later on the branch towards the Pike cluster and which are shared by Stuart and Karen's brother. We do not yet know the chronological order of the SNPs within the boxes that have been drawn. Some of the nine variants named BY36062 to BY36070 may pre-date settler James, and so they ought to be shared by all of his patrilineal descendants. But other of these nine may have arisen with the birth of James' son James Jr, in which the descendants of James Jr would carry them, but the descendants of James' other sons Jeremiah and John would not carry them. All nine of the variants that FTDNA currently lists as separating Stuart and Karen's brother are included below (bearing in mind that some of these may be filtered away as FTDNA performs updates to its protocols).
Group 2
Finally we have Group 2 which currently consists only of Pikes with known ancestry from Conception Bay in eastern Newfoundland. This is the group to which I belong, and so is one that I have a vested interest in learning about. And to whet your interest a bit, I believe that the Big Y results that we now have are revealing something truly fascinating.We have three Big Y results for Group 2, from Rodney (kit 399984), me - David (kit 23996) and from my father Angus (N21510). Although the Pikes of Conception Bay are known to have been resident at Poole in the late 1600s, most of us get stuck in our genealogy around the year 1800. That's the case for the three of us with Big Y test results.
My father and I descend from a Thomas Pike whose grandson Henry was likely born about 1809, so we reckon that Thomas was born about 1750 or so. The family lived at Carbonear, where there is a gravestone for a Thomas Pike who died in 1835 at age 100. Possibly this might be our ancestor, but as yet I am unsure.
Rodney's family also hails from Carbonear, where his ancestor Timothy Pike initially resided before moving to Channel on the southwest coast of Newfoundland, where he and his wife Patience continued to build their family. At Carbonear and Channel, Timothy and Patience had at least ten children from 1821 to 1841. We do not have a birth record or a marriage record or a death record for Timothy, so we can only guess from the years in which he was fathering children that his birthdate was around 1800 or perhaps slightly earlier.
To try to establish an upper bound on the TMRCA for Rodney on the one hand, and me and my father on the other, what is known is that in 1681 a Thomas Pike was recorded as present in Carbonear. In all likelihood this was the Thomas Pike who married Susanna Bird at Poole in Dorset in 1680, where he continued to reside. To clarify that point, although present in Newfoundland in 1681, Thomas and many others would annually sail back and forth between England and Newfoundland to partake in what was known as the migratory fishery. This fishery and the annual commuting of its workforce from one side of the Atlantic Ocean to the other carried on for the better part of the 18th century. Exactly when any particular Pikes actually chose to settle and make Carbonear their primary home is lost to time, although by the early 1800s (when the local church records commence) we find a multitude of Pike families firmly established at Carbonear.
Thomas Pike of Poole did us the favour of leaving a last will and testament, written on 20 March 1703, in which he describes himself as a mariner about to depart on a voyage to Newfoundland. He goes on to name two of his four sons and two of his three daughters. I've transcribed his will, which can be read on this webpage. Work that I've done to trace the descendants of mariner Thomas Pike has shown that several of them eventually settled in Carbonear. However, for most Pikes at Carbonear (myself included) it remains an obstacle to trace back to Thomas via an unbroken name-by-name generation-by-generation genealogy. For my own line, I think I'm close, but it's going to take some more work to be sure.
Anyway the generally held belief is that mariner Thomas Pike from Poole is the common progenitor of the Pikes at Carbonear. Possibly Rodney and I might both descend from the same son of Thomas, or even the same grandson, but in any event Thomas' birth ought to provide an upper bound on the TMRCA for Rodney versus me and my father. And with mariner Thomas having married in 1680, putting his birth around 1650 or so, we have a TMRCA of about 368 years. And based on this, as well as the estimated average of "one Big Y mutation per century per lineage" we would therefore anticipate that Rodney should be separated from me and my father by about 7 (or fewer) variants.
We've seen (albeit with very limited data) from Groups 1 and 20 that one mutation per century per lineage may be too high an estimate. Values of 0.40 and 0.45 appear to be more appropriate for the results that we have been able to explore, and if these lower values were to also apply to Group 2, then there ought to be about three (or fewer) variants separating Rodney from me and my father.
So imagine my excitement when discovering that FTDNA reports that Rodney differs from my father by 10 variants, and that he differs from me by 14. Working with the 10 differences between my father and Rodney we obtain a TMRCA of 500 years when using the estimate of "one mutation per century per lineage" and a TMRCA of more than 1000 years if mutation rates of 0.40 or 0.45 are used. Such TMRCA values imply that Rodney and I do not share mariner Thomas as a common ancestor, but instead that our most recent common ancestor pre-dates Thomas by several generations.
That Rodney's results have so many differences when compared to me and my father is something that I had actually hoped for even before the Big Y test results were reported. If you take a look at the results from the 37, 67 and 111 STR-based markers for Group 2 you will observe that Rodney has several STR marker values that are distinct from all the other men in Group 2. For instance, Rodney has the only 15 on the 9th marker, the only 16 on the 13th marker, the only 12 on the 17th marker, and so forth. Even from just looking at STR markers, somehow Rodney appears to be a bit more different from everyone else in the group than anybody else is. I was curious to see if this distinctiveness would be sustained within Rodney's Big Y results. Having now found that Rodney and my father differ by ten variants it would appear that the distinction is holding up.
But let's pause for a moment and recall our first impression of the results for Stuart and Karen's brother in Group 6 was that they too had a large difference between them. In their case a number of inconsistencies in how the variants were reported were found, and we now expect that FTDNA will revise the number of differences downward. Might something similar be taking place within Group 2? Well, maybe, but I don't think so.
Of the ten differences that are reported between Rodney and my father, seven are named variants that can be inspected with the Big Y Browser tool. And in contrast to the inconsistency that we saw happening in Group 6, this time they generally look strong and convincing. For example, here's what the browser shows for Rodney at BY23385:
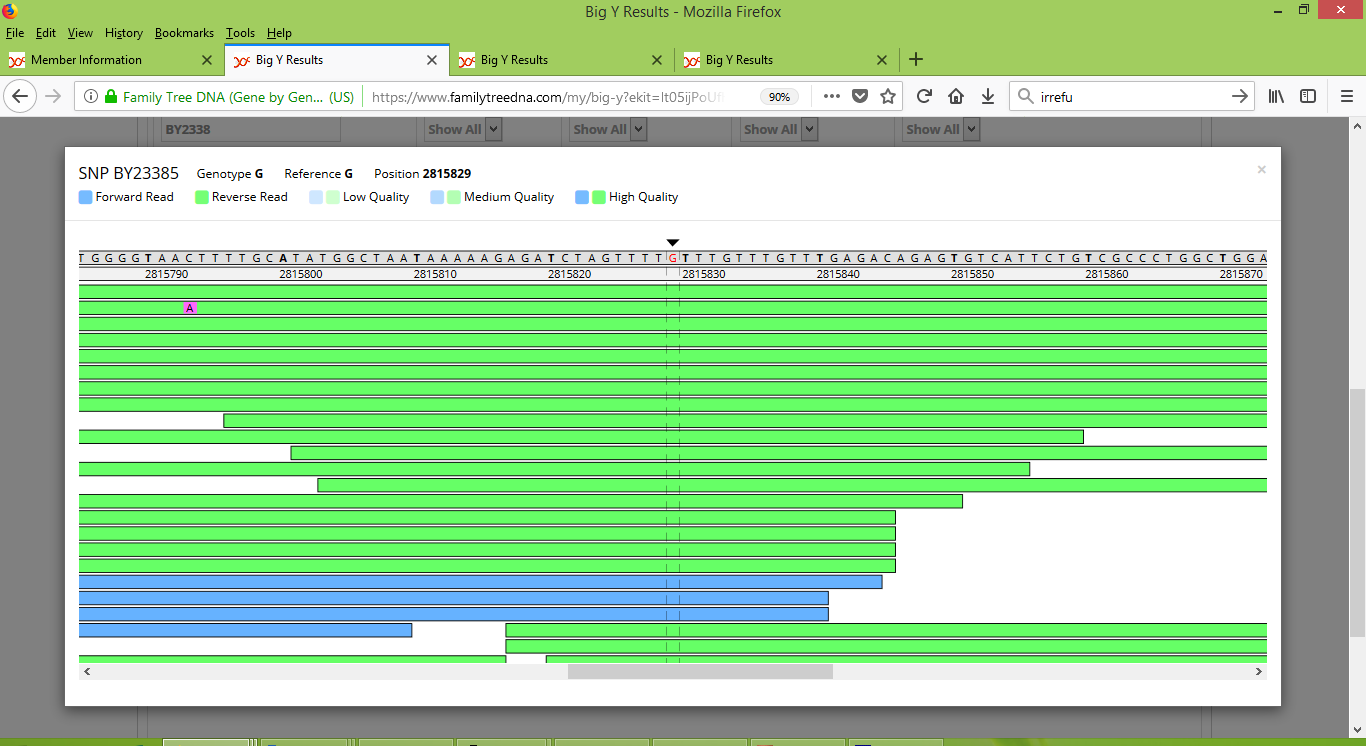
And here is what the browser shows for me at BY23385:
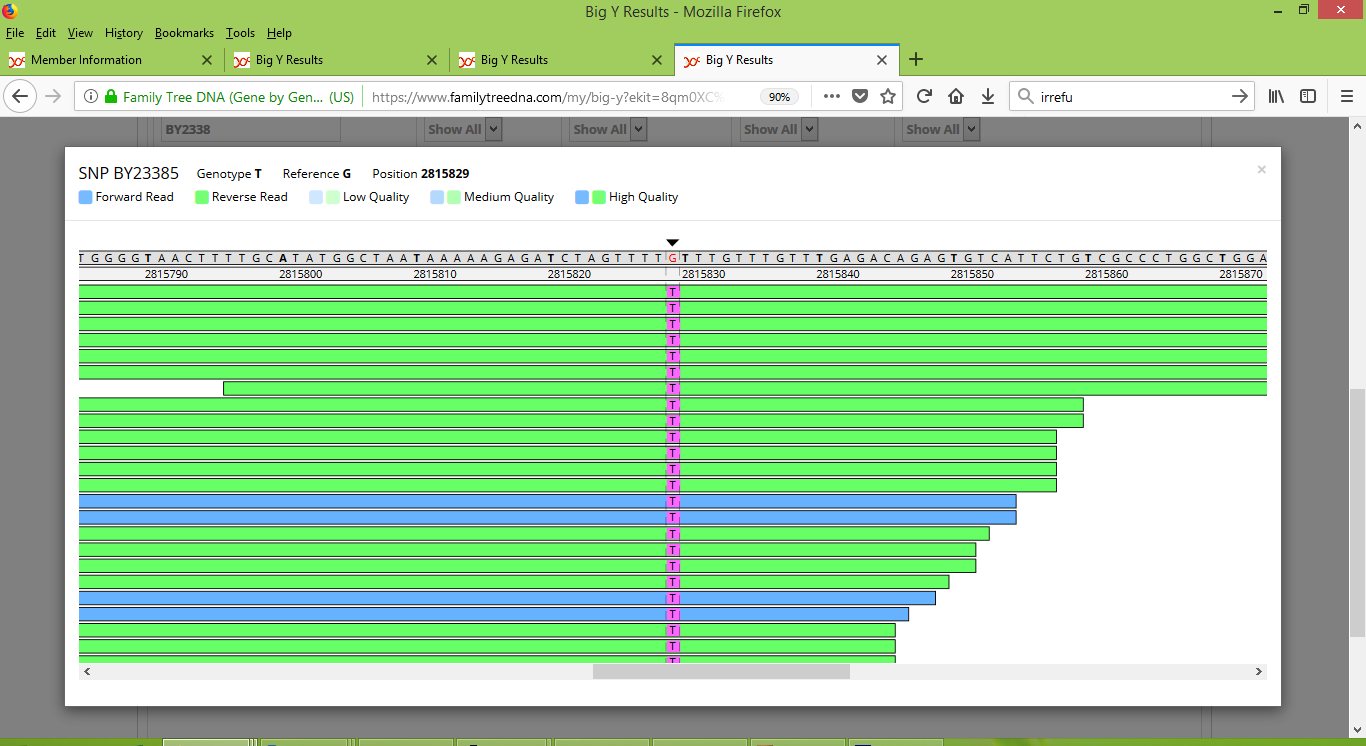
It is clearly evident that Rodney does not carry the BY23385 variant, whereas I definitely do. It is a real difference between us.
As a side question though, you may be wondering how it is that me and my father are reported as having four different variants. Three of these four are named, and so can be inspected with the Big Y Browser tool. As it happens all three of these named variants are located in close proximity and appear together in the following screenshots. This first screenshot shows my results for BY29670 and about thirteen positions to its right also BY29671 and BY29672:
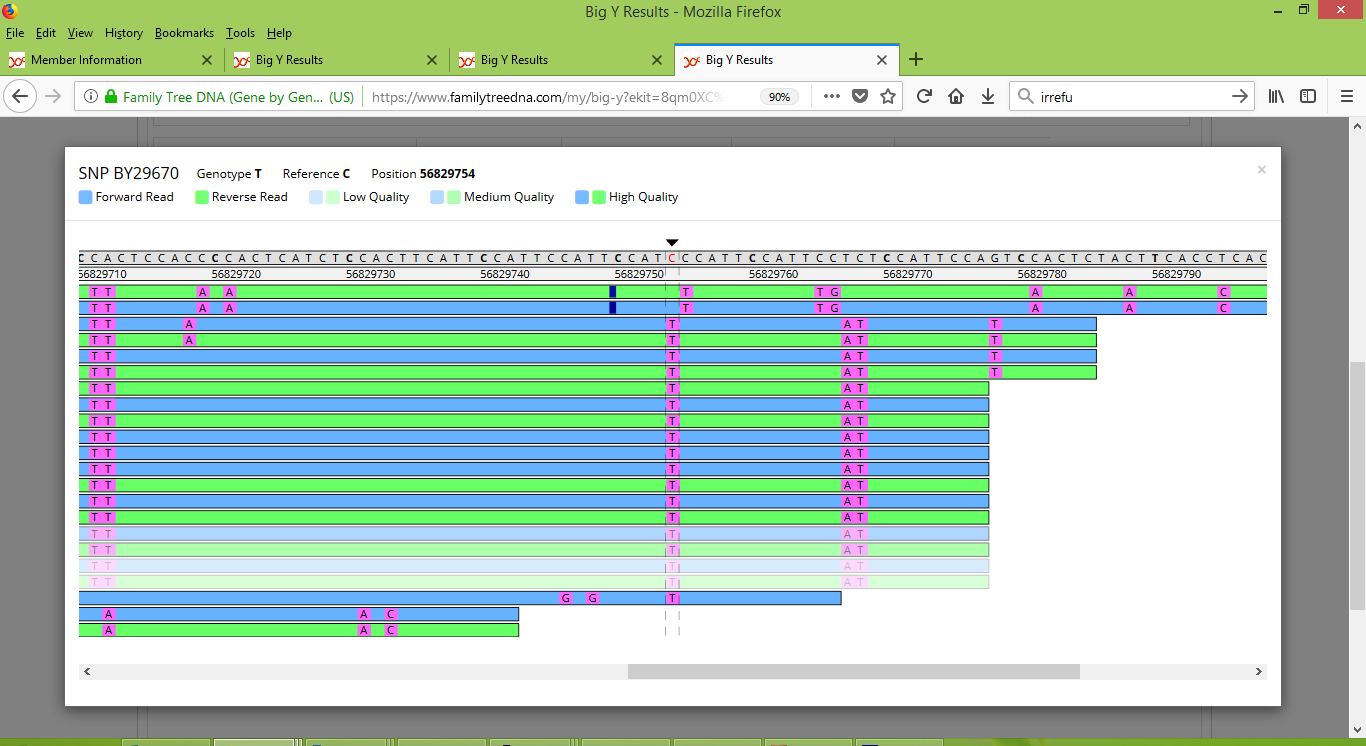
And here's a screenshot that shows my father's results for BY29670, BY29671 and BY29672:
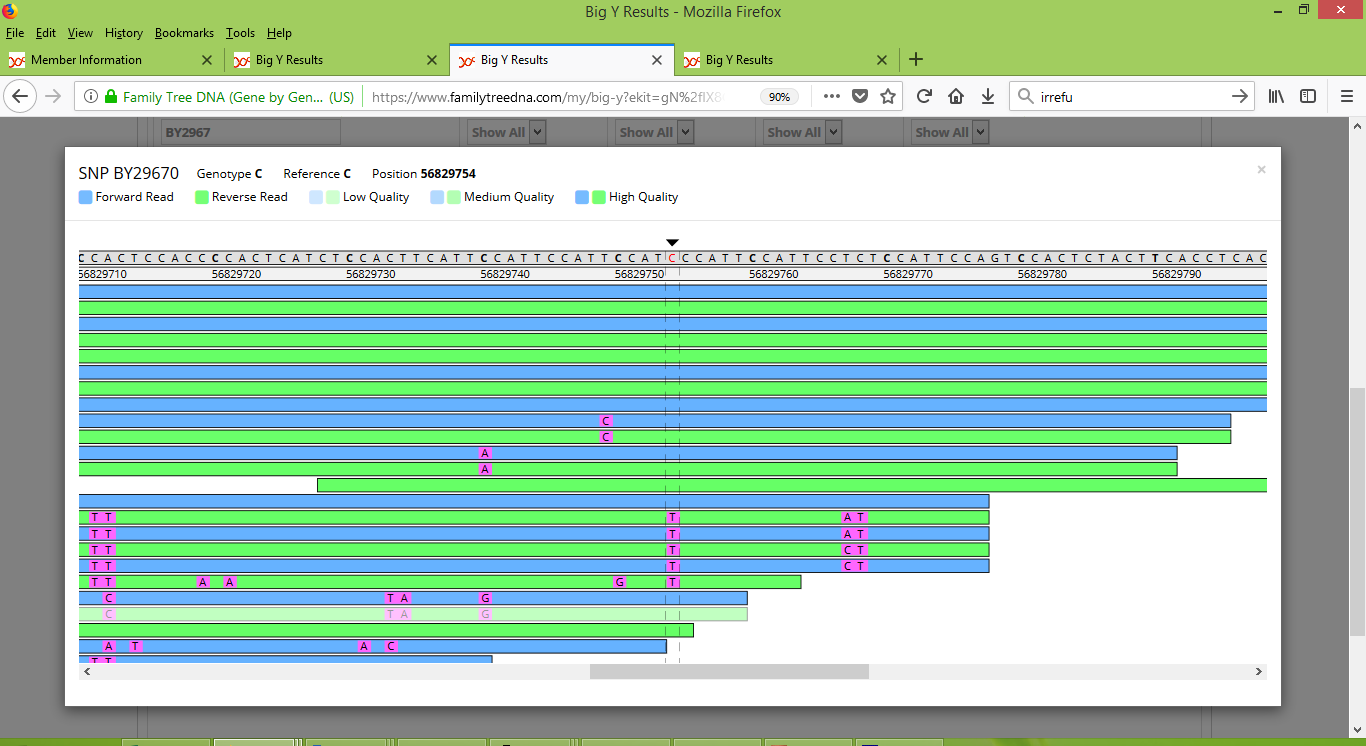
Note that for these three variants, my results appear to be consistently strong and positive. On the other hand, my father's results appear inconsistent. For a few of the scans he scored positive results but for most he scored negative. For these variants I think there's enough inconsistency to raise doubt about whether they should be counted as differences between me and my father.
But what about Rodney, how does he measure up at these three locations? These variants were among the 14 that FTDNA reported as separating me and Rodney. Below is a screenshot of Rodney's results, showing that he too is inconsistent at these three locations.
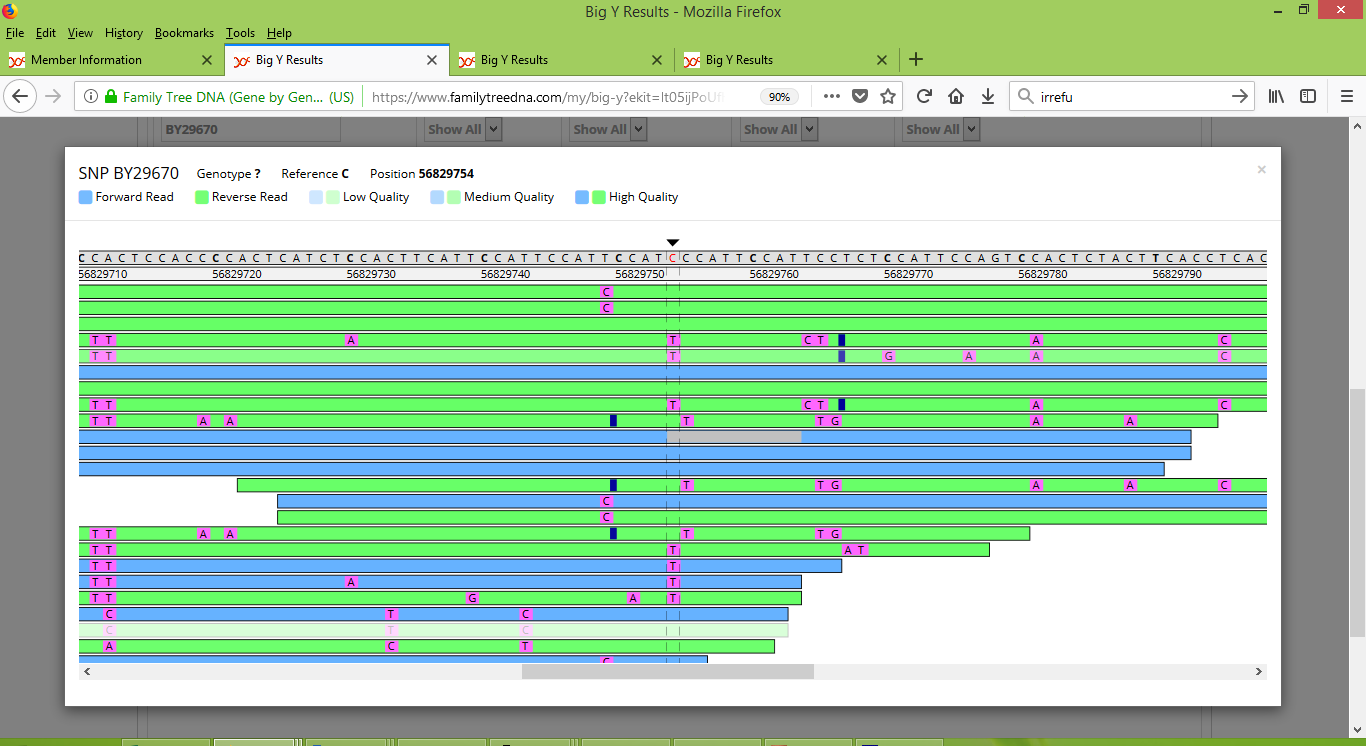
While I don't know why these three variants exhibit this inconsistency, I think that we can discount them from being considered bona fide differences between me, my father and Rodney.
Meanwhile, when looking at each of our lists of unnamed variants, FTDNA presents 14 for Rodney, 7 for my father and five for me. Two of these variants are shared by all three of us, and a further four are shared by Rodney and my father (probably I share these four too, but I might have No Calls at these locations). These shared variants will become named in due course. But that still leaves a number of additional variants that aren't shared and which are not yet listed among the 10 or 14 variants that reportedly separate Rodney from my father and me. With further analysis these could potentially be deemed to be differences, which would further increase the separation between us.
The conclusion that I'm drawing from the results on hand for Group 2 is that there really is a significant difference between Rodney's results and those of me and my father. And that it is enough to indicate that our most recent common ancestor lived sometime before mariner Thomas of Poole. This is a fascinating revelation, in part because it indicates that the Pike family of Group 2 goes back even further than we've been able to knowingly trace it in historical records. And with such an earlier origin, there would have been ample opportunity for other offshoots of the family to have arisen and still be out there somewhere (by which I mean distant cousins whose ancestors might have never been engaged with the migratory fishery to Newfoundland). The profound lack of genetic matches from places other than Newfoundland is therefore something of a puzzle for those of us in Group 2. If we have distant Pike relatives out there, where are they?
Another impressive insight that we can draw from these results is that although Rodney's line and mine may have diverged from a common ancestor who lived well before the time of mariner Thomas at Poole, these two lines nevertheless remained in close proximity to one another, including both making their way to the town of Carbonear in Newfoundland. Given the STR-based results that we have for Group 2, it would appear that Rodney's line may not have been as prolific or populous. However, there are still several Pike lines from Carbonear and elsewhere in Newfoundland who have not engaged in Y-DNA testing, so we may yet find some more representatives from Rodney's side of the family tree. Or it may be that Rodney's ancestor Timothy was the last of his line at Carbonear, and when he moved away nobody was left behind who shared the distinctive markers within his Y-DNA. We can easily come up with more questions, but for answers we will have to be patient and wait for more DNA results to come forward.
Let's now turn our attention toward the SNP-based tree for Group 2. Everybody in this group fits into the large R1b Haplogroup and in particular to a subgroup that is defined by the L21 variant, then into the even smaller (but still large) Z253 subroup, and then into its L554 subgroup. The L554 subgroup is relatively small. Since its discovery in March of 2011 only a handful of surname clusters have been found within it, namely Humphrey, Pike and Wilcox, along with some sporadic instances of men who don't fit into any surname clusters. About a year ago one each of Humphrey, Pike and Wilcox had done the Big Y test, which has enabled the age of the L554 subgroup to be estimated at upwards of 2000 years.
Although FTDNA does not list any non-Pike matches for any of me, my father or Rodney, the Big Y results from Humphrey and Wilcox help to define which variants pre-date our surnames (similar to the situation with Schupp in Group 6). Here then is the SNP-based tree for Group 2, without the three inconsistent variants that FTDNA initially suggested as different for me and my father.
Note that from the point at which Rodney's line branches away from mine, his line appears to have accumulated only two mutations, whereas the line to my father appears to have acquired eight. That might make sense if Rodney's line happened to have a sequence of men who were fathering children when they were advanced in years, and if my line had men who became fathers when they were relatively young (i.e., if my line has gone through more generations than Rodney's). What I think is actually happening though is that where both my father and I have done the Big Y test, FTDNA has had the benefit of two men's results to validate our variants and confirm their relevance. On the other hand, the several unnamed variants that Rodney carries but which aren't yet reported as differences with me and my father have not yet had the benefit of this kind of second opinion obtained from another person's results (that is, another person who carries many of the same variants as Rodney). Once the variants in Rodney's results get validated in this way I would imagine that FTDNA will report additional differences, stemming from additional mutations along Rodney's patriline. If and when that happens, then that would further increase the number of differences between Rodney and my father, beyond the current 10, and would in turn reinforce the theory that our MRCA lived sometime prior to mariner Thomas Pike of Poole.
The collection of seven variants that Rodney, me and my father all share, but which aren't shared by the non-Pike that is listed, are simply listed in alphanumeric order. We do not yet know their chronological order. Some may pre-date the Pike surname for Group 2, while others may have arisen after the surname came into use (but before the point in the family tree at which Rodney's line split from mine). The eight variants that my father has but Rodney lacks are likewise not presented in chronological order. As more members of Group 2 take the Big Y test we will be better able to position these variants in relation to one another, and at the same time develop the structure of our family tree. Said another way, we can rely on our genetic variants to piece together our family tree and put the various branches onto it, even if we don't have complete historical records. And since nearly all of us in Group 2 get genealogically stuck around the year 1800, this is simply wonderful. It is the kind of thing that I've wanted to accomplish since the Pike DNA Project came into being back in 2004. This is truly exciting!
Lessons Learned and What's Next
First of all, I have to say that I am impressed with the power of the Big Y test and what we may be able to discern from these Big Y results. The corresponding implications are giving us brand new insights into our history that I don't think we could have figured out any other way.I should emphasise that like all genealogical theories, the conclusions that we have drawn are based only on the available information. As new information becomes available, its interpretation may result in revisions and adapted conclusions. For instance, we may find that in some of our project's genetic clusters the mutation rate is low (such as 0.40 mutations per century per line) but it may be found to be high in others. Ultimately our goal is to determine the truth and to better understand our family history.
One thing that I hope was evident from the foregoing discussion and examples is that this is very much a group effort. It is only by working together and having DNA reults from multiple people that we will be able to take full advantage of what genetic genealogy has to offer for us. Even if you may not gain substantial or immediate benefit from your own results, your results may nevertheless hold a key to solving part of the puzzle that we are collectively putting back together.
I want to encourage as many people as possible to do the Big Y test, most especially for those lines that haven't yet been tested. To name a few such lines, in Group 1 we do not yet have any Big Y results from people who do not descend from settler John Pike, such as those who remained behind when he left England and/or those who carry a marker value of 24 on the second of the 111 STR-based markers. Big Y results from such people would help to determine which Big Y variants pre-date the Pike surname versus which ones are shared by all of Group 1. And as I've tried to do in my analysis of Group 2, a broad set of results should enable us to estimate how old the family is... that is, how long it has been since the family's MRCA. Meanwhile, additional Big Y results from descendants of settler John will help us to map out which variants belong to which specific branches of the family. Additional results may also help to validate unnamed variants that FTDNA isn't yet reporting as differences.
Much the same can be said for our other genetic groups as well, so I'll just echo the point that it will be good to get as many test results to work with as we can. Moreover, by mapping out the SNP-based tree for each of our family groups, it should become much easier to determine where people with incomplete pedigrees and brickwalls belong.
However, keep in mind that Big Y tests should be treated as exploratory research that may not always provide information of genealogical significance in the near term (so please don't set your expectations too high).
As it pertains to my desire to encourage more people to upgrade and get tested, I will briefly mention that our project has so far received $30 in donations to help support Big Y testing. This particular donation is aimed towards our Group 1.
An Academic Research Project in England
Finally, for those who have taken (or plan to take) the Big Y test, I recently became aware of an academic research project being carried out by investigators at the University of Leicester. They appear to have a goal of further developing the human Y-chromosome SNP tree, and are especially interested in families that originate in the southwest of England (which is indeed the case for many Pike lines). As I haven't yet heard directly from the investigators, I don't yet have much more that I can say about their project. For more information on their project, please refer to their webpage.
Last Modified: Wednesday, 28 March 2018, 17:18:11 NDT